22 GLMM, Proportion Outcome: Meta Analysis (Hox ch 6)
See Appendix E of the Joop Hox text.
https://uoepsy.github.io/msmr/2122/lectures/msmr_lec02_LogisticMLM.html#1
https://commresearch.arizona.edu/classes/comm640/640_Book/docs/multilevel-logistic-regression.html
https://stats.stackexchange.com/questions/544937/when-is-it-appropriate-to-set-nagq-0-in-glmer
https://bookdown.org/animestina/phd_july_19/now-for-advanced-logistic-mixed-effects.html
22.1 PREPARATION
22.1.2 Background
NESTING
* source study id on which multiple conditions were reported (most studies reported on 2 of the 3 modes)
Study Characteristics:
respisrrtype of study reported proportionsCrcompletion rate, # respondents / total sample sizeRrresponse rate, sample size was corrected for sampling frame errorsyear0 = 1947, the oldest yearsaliencysaliency of the questionnaire, 0 = not at all, 1 = somewhat, 2 = highly
Condition Characteristics:
modedta collection method, 3-levels: Face-to-face, Telephone, or Mailteldummy code for mode = Telephonemaildummy code for mode = Mailresponseeither the completion rate or response ratedenomnumber of respondents approached
22.1.3 Read in the data
df_raw <- haven::read_sav("data/metaresp.sav") %>%
haven::as_factor() %>%
haven::zap_label() %>%
haven::zap_formats() %>%
haven::zap_widths() Rows: 105
Columns: 14
$ source <dbl> 1, 1, 1, 2, 2, 2, 3, 3, 4, 4, 5, 5, 6, 6, 7, 7, 8, 8, 9, 9, 9…
$ mode <fct> ftf, tel, mail, ftf, tel, mail, ftf, mail, ftf, tel, ftf, tel…
$ cr <dbl> 0.60, 0.64, 0.49, NA, NA, NA, 0.69, 0.72, 0.80, 0.82, 0.32, 0…
$ rr <dbl> NA, NA, NA, 0.60, 0.52, 0.60, 0.71, 0.73, 0.88, 0.86, 0.36, 0…
$ ftfdum <fct> ftf condition, not ftf, not ftf, ftf condition, not ftf, not …
$ teldum <fct> not telephone, telephone condition, not telephone, not teleph…
$ maildum <fct> notmail, notmail, mail condition, notmail, notmail, mail cond…
$ year <dbl> 25, 25, 25, 35, 35, 35, 34, 34, 35, 35, 13, 13, 16, 16, 22, 2…
$ saliency <fct> some, some, some, not, not, not, high, high, some, some, not,…
$ denomtot <dbl> 267, 159, 320, NA, NA, NA, 125, 118, 296, 374, 306, 374, 481,…
$ denomcor <dbl> NA, NA, NA, 220, 229, 893, 121, 116, 269, 357, 272, 323, NA, …
$ response <dbl> 0.60, 0.64, 0.49, 0.60, 0.52, 0.60, 0.71, 0.73, 0.88, 0.86, 0…
$ denom <dbl> 267, 159, 320, 220, 229, 893, 121, 116, 269, 357, 272, 323, 4…
$ respisrr <fct> cr, cr, cr, rr, rr, rr, rr, rr, rr, rr, rr, rr, cr, cr, cr, c…22.1.4 Long Format
One line per mode per source
df_long <- df_raw %>%
dplyr::rename_all(tolower) %>%
dplyr::mutate_if(is.factor, ~forcats::fct_relabel(.x, stringr::str_to_title)) %>%
dplyr::mutate(res = round(response*denom, 0)) %>%
dplyr::mutate(tel = as.numeric(mode == "Tel"))%>%
dplyr::mutate(mail = as.numeric(mode == "Mail")) %>%
dplyr::mutate(yr = year + 1947) %>%
dplyr::mutate(yr47 = year) %>%
dplyr::mutate(yr77 = year - 30) %>%
dplyr::select(source,
yr, yr47, yr77,
saliency,
mode, tel, mail,
respisrr,
response, res, denom)Rows: 105
Columns: 12
$ source <dbl> 1, 1, 1, 2, 2, 2, 3, 3, 4, 4, 5, 5, 6, 6, 7, 7, 8, 8, 9, 9, 9…
$ yr <dbl> 1972, 1972, 1972, 1982, 1982, 1982, 1981, 1981, 1982, 1982, 1…
$ yr47 <dbl> 25, 25, 25, 35, 35, 35, 34, 34, 35, 35, 13, 13, 16, 16, 22, 2…
$ yr77 <dbl> -5, -5, -5, 5, 5, 5, 4, 4, 5, 5, -17, -17, -14, -14, -8, -8, …
$ saliency <fct> Some, Some, Some, Not, Not, Not, High, High, Some, Some, Not,…
$ mode <fct> Ftf, Tel, Mail, Ftf, Tel, Mail, Ftf, Mail, Ftf, Tel, Ftf, Tel…
$ tel <dbl> 0, 1, 0, 0, 1, 0, 0, 0, 0, 1, 0, 1, 0, 0, 0, 1, 0, 1, 0, 1, 0…
$ mail <dbl> 0, 0, 1, 0, 0, 1, 0, 1, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1…
$ respisrr <fct> Cr, Cr, Cr, Rr, Rr, Rr, Rr, Rr, Rr, Rr, Rr, Rr, Cr, Cr, Cr, C…
$ response <dbl> 0.60, 0.64, 0.49, 0.60, 0.52, 0.60, 0.71, 0.73, 0.88, 0.86, 0…
$ res <dbl> 160, 102, 157, 132, 119, 536, 86, 85, 237, 307, 98, 71, 462, …
$ denom <dbl> 267, 159, 320, 220, 229, 893, 121, 116, 269, 357, 272, 323, 4…df_long %>%
dplyr::select(source, respisrr, mode, tel, mail, yr, saliency, response, denom, res) %>%
psych::headTail(top = 10) %>%
flextable::flextable() %>%
apaSupp::theme_apa(caption = "Meta Data for Response Rate, Long Format")source | respisrr | mode | tel | yr | saliency | response | denom | res | |
|---|---|---|---|---|---|---|---|---|---|
1 | Cr | Ftf | 0 | 0 | 1972 | Some | 0.6 | 267 | 160 |
1 | Cr | Tel | 1 | 0 | 1972 | Some | 0.64 | 159 | 102 |
1 | Cr | 0 | 1 | 1972 | Some | 0.49 | 320 | 157 | |
2 | Rr | Ftf | 0 | 0 | 1982 | Not | 0.6 | 220 | 132 |
2 | Rr | Tel | 1 | 0 | 1982 | Not | 0.52 | 229 | 119 |
2 | Rr | 0 | 1 | 1982 | Not | 0.6 | 893 | 536 | |
3 | Rr | Ftf | 0 | 0 | 1981 | High | 0.71 | 121 | 86 |
3 | Rr | 0 | 1 | 1981 | High | 0.73 | 116 | 85 | |
4 | Rr | Ftf | 0 | 0 | 1982 | Some | 0.88 | 269 | 237 |
4 | Rr | Tel | 1 | 0 | 1982 | Some | 0.86 | 357 | 307 |
... | ... | ... | ... | ... | ... | ... | |||
77 | Rr | 0 | 1 | 1992 | Some | 0.75 | 92 | 69 | |
78 | Rr | Ftf | 0 | 0 | 1992 | Some | 0.51 | 476 | 243 |
78 | Rr | Tel | 1 | 0 | 1992 | Some | 0.66 | 406 | 268 |
78 | Rr | 0 | 1 | 1992 | Some | 0.67 | 372 | 249 |
22.1.5 Wide Format
One line per source
df_wide <- df_long %>%
dplyr::group_by(source) %>%
dplyr::mutate(n_modes = n()) %>%
dplyr::mutate(respisrr = dplyr::first(respisrr)) %>%
dplyr::ungroup() %>%
dplyr::select(source, n_modes, yr, yr47, yr77, respisrr, saliency, mode, response) %>%
tidyr::pivot_wider(names_from = mode,
names_prefix = "resp_",
values_from = response)Rows: 48
Columns: 10
$ source <dbl> 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 14, 16, 17, 18, 19, 2…
$ n_modes <int> 3, 3, 2, 2, 2, 2, 2, 2, 3, 2, 2, 2, 3, 3, 3, 2, 2, 2, 2, 3, …
$ yr <dbl> 1972, 1982, 1981, 1982, 1960, 1963, 1969, 1969, 1984, 1947, …
$ yr47 <dbl> 25, 35, 34, 35, 13, 16, 22, 22, 37, 0, 31, 31, 31, 20, 20, 2…
$ yr77 <dbl> -5, 5, 4, 5, -17, -14, -8, -8, 7, -30, 1, 1, 1, -10, -10, -6…
$ respisrr <fct> Cr, Rr, Rr, Rr, Rr, Cr, Cr, Cr, Rr, Cr, Rr, Rr, Rr, Rr, Rr, …
$ saliency <fct> Some, Not, High, Some, Not, Some, High, High, Some, High, So…
$ resp_Ftf <dbl> 0.60, 0.60, 0.71, 0.88, 0.36, 0.96, 0.64, 0.68, 0.71, 0.97, …
$ resp_Tel <dbl> 0.64, 0.52, NA, 0.86, 0.22, NA, 0.72, 0.57, 0.73, NA, 0.59, …
$ resp_Mail <dbl> 0.49, 0.60, 0.73, NA, NA, 0.92, NA, NA, 0.66, 0.82, NA, NA, …df_wide %>%
dplyr::select(-yr47, -yr77) %>%
psych::headTail(top = 10) %>%
flextable::flextable() %>%
apaSupp::theme_apa(caption = "Meta Data for Response Rate, Wide Format")source | n_modes | yr | respisrr | saliency | resp_Ftf | resp_Tel | resp_Mail |
|---|---|---|---|---|---|---|---|
1 | 3 | 1972 | Cr | Some | 0.6 | 0.64 | 0.49 |
2 | 3 | 1982 | Rr | Not | 0.6 | 0.52 | 0.6 |
3 | 2 | 1981 | Rr | High | 0.71 | 0.73 | |
4 | 2 | 1982 | Rr | Some | 0.88 | 0.86 | |
5 | 2 | 1960 | Rr | Not | 0.36 | 0.22 | |
6 | 2 | 1963 | Cr | Some | 0.96 | 0.92 | |
7 | 2 | 1969 | Cr | High | 0.64 | 0.72 | |
8 | 2 | 1969 | Cr | High | 0.68 | 0.57 | |
9 | 3 | 1984 | Rr | Some | 0.71 | 0.73 | 0.66 |
10 | 2 | 1947 | Cr | High | 0.97 | 0.82 | |
... | ... | ... | ... | ... | ... | ||
75 | 2 | 1987 | Rr | Some | 0.79 | 0.86 | |
76 | 2 | 1983 | Cr | Not | 0.64 | 0.54 | |
77 | 3 | 1992 | Rr | Some | 0.54 | 0.64 | 0.75 |
78 | 3 | 1992 | Rr | Some | 0.51 | 0.66 | 0.67 |
22.2 EXPLORATORY DATA ANLAYSIS
22.2.2 Missing Data - None!
df_long %>%
VIM::aggr(numbers = TRUE, # shows the number to the far right
prop = FALSE) # shows counts instead of proportions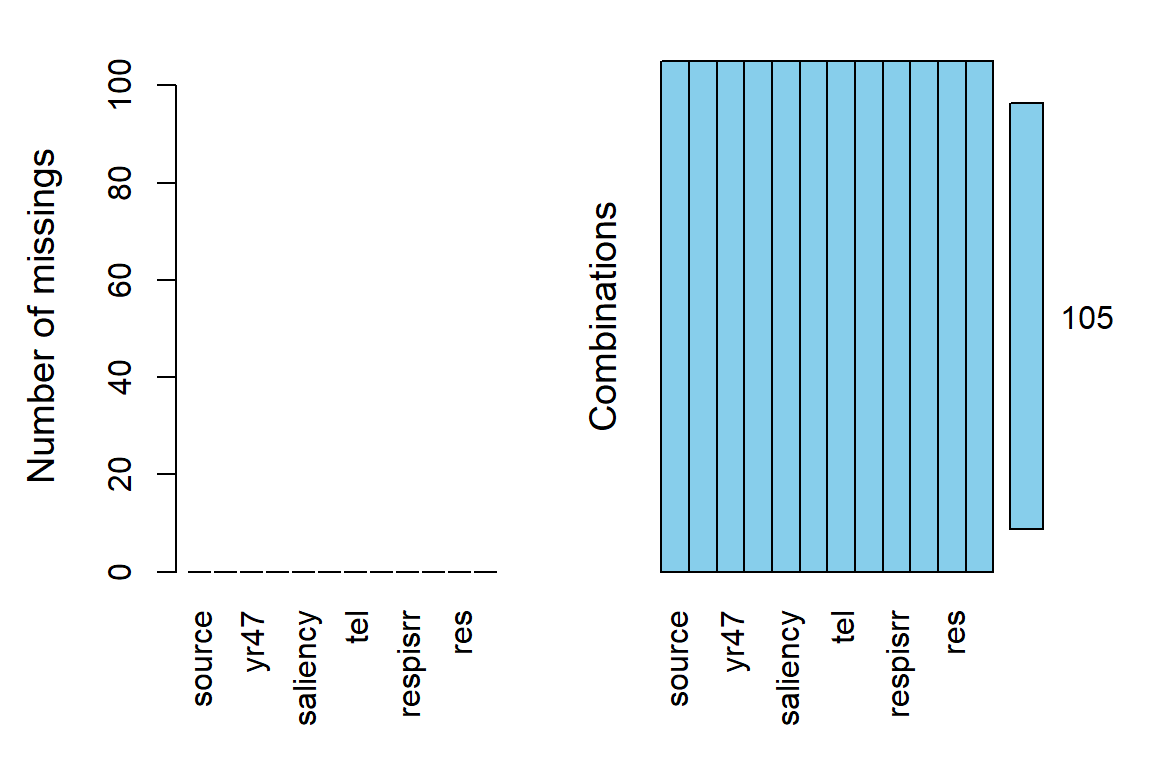
22.2.3 Descriptive Summary
df_wide %>%
dplyr::select(n_modes,
yr,
resp_Ftf,
resp_Tel,
resp_Mail) %>%
apaSupp::tab_desc(caption="Descriptive Summary of Sources")NA | M | SD | min | Q1 | Mdn | Q3 | max | |
|---|---|---|---|---|---|---|---|---|
n_modes | 0 | 2.19 | 0.53 | 1.00 | 2.00 | 2.00 | 2.25 | 3.00 |
yr | 0 | 1,976.06 | 9.77 | 1,947.00 | 1,969.00 | 1,978.00 | 1,983.00 | 1,992.00 |
resp_Ftf | 8 | 0.75 | 0.14 | 0.36 | 0.65 | 0.73 | 0.87 | 0.99 |
resp_Tel | 11 | 0.70 | 0.16 | 0.22 | 0.59 | 0.73 | 0.81 | 0.99 |
resp_Mail | 20 | 0.65 | 0.18 | 0.19 | 0.63 | 0.69 | 0.75 | 0.95 |
Note. N = 48. NA = not available or missing; Mdn = median; Q1 = 25th percentile; Q3 = 75th percentile. | ||||||||
df_long %>%
dplyr::select(mode,
yr, saliency,
respisrr,
response, res, denom) %>%
apaSupp::tab1(split = "mode")
| Total | Ftf | Tel | Mail | p-value |
|---|---|---|---|---|---|
yr | 1,976.74 (9.60) | 1,976.73 (10.35) | 1,977.68 (8.64) | 1,975.54 (9.88) | .662 |
saliency | .683 | ||||
Not | 19 (18.1%) | 6 (15.0%) | 9 (24.3%) | 4 (14.3%) | |
Some | 60 (57.1%) | 22 (55.0%) | 20 (54.1%) | 18 (64.3%) | |
High | 26 (24.8%) | 12 (30.0%) | 8 (21.6%) | 6 (21.4%) | |
respisrr | .932 | ||||
Cr | 39 (37.1%) | 14 (35.0%) | 14 (37.8%) | 11 (39.3%) | |
Rr | 66 (62.9%) | 26 (65.0%) | 23 (62.2%) | 17 (60.7%) | |
response | 0.71 (0.16) | 0.75 (0.14) | 0.70 (0.16) | 0.65 (0.18) | .068 |
res | 768.32 (2,095.35) | 1,216.85 (3,213.75) | 566.76 (1,016.13) | 393.93 (342.48) | .198 |
denom | 1,003.32 (2,222.26) | 1,458.15 (3,365.10) | 772.97 (1,163.73) | 657.96 (597.34) | .327 |
Note. Continuous variables are summarized with means (SD) and significant group differences assessed via independent one-way analysis of vaiance (ANOVA). Categorical variables are summarized with counts (%) and significant group differences assessed via Chi-squared tests for independence. | |||||
* p < .05. ** p < .01. *** p < .001. | |||||
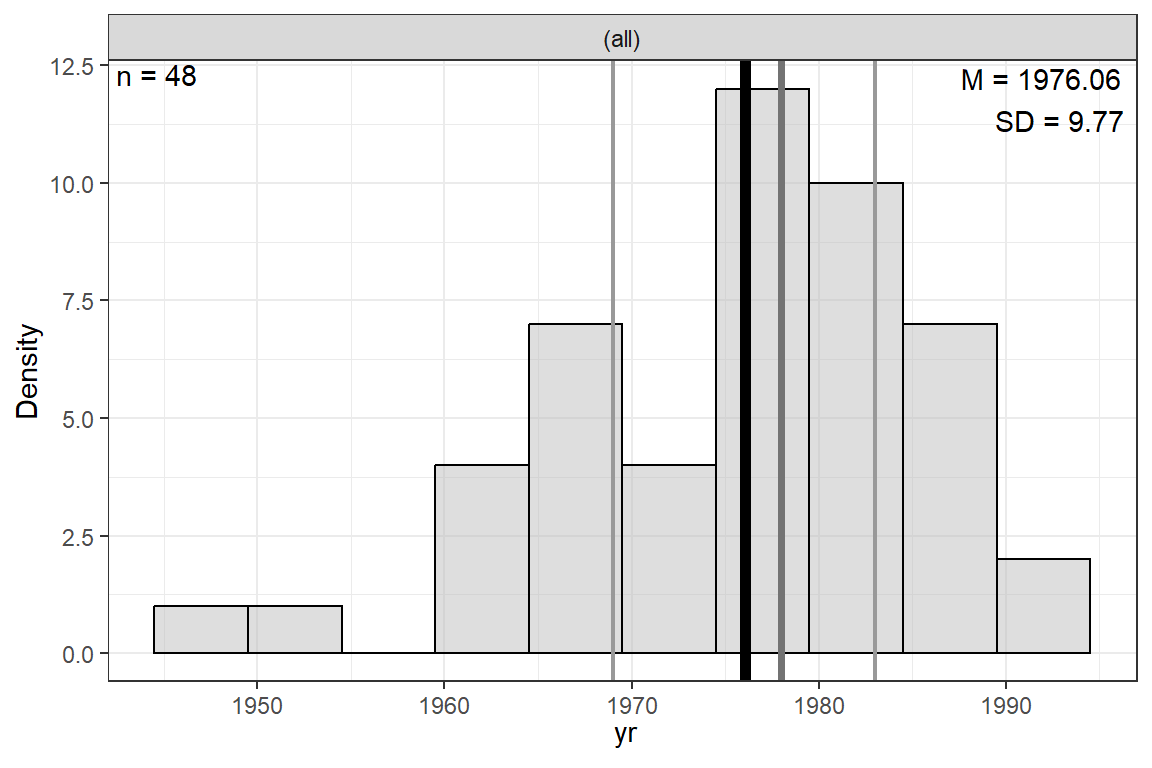
Figure 22.1
Distribution of Source Year (0 for 1947)
22.3 MULTILEVEL MODELING
22.3.1 Base Model
22.3.1.1 PQL
fit_glmer_0 <- lme4::glmer(cbind(res, denom - res) ~ respisrr + (1|source),
data = df_long,
family = binomial,
nAGQ = 0)Generalized linear mixed model fit by maximum likelihood (Adaptive
Gauss-Hermite Quadrature, nAGQ = 0) [glmerMod]
Family: binomial ( logit )
Formula: cbind(res, denom - res) ~ respisrr + (1 | source)
Data: df_long
AIC BIC logLik -2*log(L) df.resid
2236 2244 -1115 2230 102
Scaled residuals:
Min 1Q Median 3Q Max
-11.973 -1.893 0.062 1.881 11.112
Random effects:
Groups Name Variance Std.Dev.
source (Intercept) 0.92 0.959
Number of obs: 105, groups: source, 48
Fixed effects:
Estimate Std. Error z value Pr(>|z|)
(Intercept) 0.5907 0.1440 4.1 4.1e-05 ***
respisrrRr 0.7012 0.0579 12.1 < 2e-16 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Correlation of Fixed Effects:
(Intr)
respisrrRr -0.24322.3.1.2 Laplace
fit_glmer_1 <- lme4::glmer(cbind(res, denom - res) ~ respisrr + (1|source),
data = df_long,
family = binomial)Generalized linear mixed model fit by maximum likelihood (Laplace
Approximation) [glmerMod]
Family: binomial ( logit )
Formula: cbind(res, denom - res) ~ respisrr + (1 | source)
Data: df_long
AIC BIC logLik -2*log(L) df.resid
2236 2244 -1115 2230 102
Scaled residuals:
Min 1Q Median 3Q Max
-11.973 -1.893 0.062 1.881 11.111
Random effects:
Groups Name Variance Std.Dev.
source (Intercept) 0.92 0.959
Number of obs: 105, groups: source, 48
Fixed effects:
Estimate Std. Error z value Pr(>|z|)
(Intercept) 0.5948 0.1440 4.13 3.6e-05 ***
respisrrRr 0.7012 0.0582 12.06 < 2e-16 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Correlation of Fixed Effects:
(Intr)
respisrrRr -0.24122.3.1.3 Compare
| PQL | Laplace | ||||
|---|---|---|---|---|---|---|
Variable | b | (SE) | p | b | (SE) | p |
(Intercept) | 0.59 | (0.14) | < .001*** | 0.59 | (0.14) | < .001*** |
respisrr | ||||||
Cr | — | — | — | — | ||
Rr | 0.70 | (0.06) | < .001*** | 0.70 | (0.06) | < .001*** |
source.var__(Intercept) | 0.92 | 0.92 | ||||
AIC | 2,236 | 2,236 | ||||
BIC | 2,244 | 2,244 | ||||
Log-likelihood | -1,115 | -1,115 | ||||
Note. | ||||||
* p < .05. ** p < .01. *** p < .001. | ||||||
22.3.2 Main Effect of Model
22.3.2.1 Random Intercepts
fit_glmer_2 <- lme4::glmer(cbind(res, denom - res) ~ respisrr + mode + (1|source),
data = df_long,
family = binomial)Generalized linear mixed model fit by maximum likelihood (Laplace
Approximation) [glmerMod]
Family: binomial ( logit )
Formula: cbind(res, denom - res) ~ respisrr + mode + (1 | source)
Data: df_long
AIC BIC logLik -2*log(L) df.resid
1939 1952 -964 1929 100
Scaled residuals:
Min 1Q Median 3Q Max
-10.278 -1.757 0.062 1.811 8.039
Random effects:
Groups Name Variance Std.Dev.
source (Intercept) 0.849 0.921
Number of obs: 105, groups: source, 48
Fixed effects:
Estimate Std. Error z value Pr(>|z|)
(Intercept) 0.9045 0.1405 6.44 1.2e-10 ***
respisrrRr 0.5274 0.0608 8.68 < 2e-16 ***
modeTel -0.1600 0.0211 -7.59 3.2e-14 ***
modeMail -0.4875 0.0282 -17.27 < 2e-16 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Correlation of Fixed Effects:
(Intr) rspsrR modeTl
respisrrRr -0.280
modeTel -0.142 0.297
modeMail -0.118 0.135 0.33522.3.2.2 Add Random Slopes
fit_glmer_3 <- lme4::glmer(cbind(res, denom - res) ~ respisrr + mode +
(1|source) + (0 + tel|source) + (0+ mail|source),
data = df_long,
family = binomial)Generalized linear mixed model fit by maximum likelihood (Laplace
Approximation) [glmerMod]
Family: binomial ( logit )
Formula: cbind(res, denom - res) ~ respisrr + mode + (1 | source) + (0 +
tel | source) + (0 + mail | source)
Data: df_long
AIC BIC logLik -2*log(L) df.resid
1150 1169 -568 1136 98
Scaled residuals:
Min 1Q Median 3Q Max
-1.2636 -0.1418 0.0099 0.1968 1.2858
Random effects:
Groups Name Variance Std.Dev.
source (Intercept) 0.766 0.875
source.1 tel 0.241 0.491
source.2 mail 0.443 0.666
Number of obs: 105, groups: source, 48
Fixed effects:
Estimate Std. Error z value Pr(>|z|)
(Intercept) 1.1429 0.2030 5.63 1.8e-08 ***
respisrrRr 0.1886 0.2346 0.80 0.42156
modeTel -0.1756 0.0926 -1.90 0.05796 .
modeMail -0.5103 0.1444 -3.53 0.00041 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Correlation of Fixed Effects:
(Intr) rspsrR modeTl
respisrrRr -0.755
modeTel -0.138 0.103
modeMail -0.174 0.120 0.13022.3.2.3 Compare
| RI | RIAS | ||||
|---|---|---|---|---|---|---|
Variable | b | (SE) | p | b | (SE) | p |
(Intercept) | 0.90 | (0.14) | < .001*** | 1.14 | (0.20) | < .001*** |
respisrr | ||||||
Cr | — | — | — | — | ||
Rr | 0.53 | (0.06) | < .001*** | 0.19 | (0.23) | .422 |
mode | ||||||
Ftf | — | — | — | — | ||
Tel | -0.16 | (0.02) | < .001*** | -0.18 | (0.09) | .058 |
-0.49 | (0.03) | < .001*** | -0.51 | (0.14) | < .001*** | |
source.var__(Intercept) | 0.85 | 0.77 | ||||
source.1.var__tel | 0.24 | |||||
source.2.var__mail | 0.44 | |||||
AIC | 1,939 | 1,150 | ||||
BIC | 1,952 | 1,169 | ||||
Log-likelihood | -964 | -568 | ||||
Note. | ||||||
* p < .05. ** p < .01. *** p < .001. | ||||||
Model | AIC | BIC | RMSE |
| |
|---|---|---|---|---|---|
Conditional | Marginal | ||||
RI | 1938.61 | 1951.88 | 0.07 | .997 | .112 |
RIAS | 1150.49 | 1169.07 | 0.01 | .996 | .062 |
Note. Larger values indicated better performance. Smaller values indicated better performance for Akaike's Information Criteria (AIC), Bayesian information criteria (BIC), and Root Mean Squared Error (RMSE). | |||||
Data: df_long
Models:
fit_glmer_2: cbind(res, denom - res) ~ respisrr + mode + (1 | source)
fit_glmer_3: cbind(res, denom - res) ~ respisrr + mode + (1 | source) + (0 + tel | source) + (0 + mail | source)
npar AIC BIC logLik -2*log(L) Chisq Df Pr(>Chisq)
fit_glmer_2 5 1939 1952 -964 1929
fit_glmer_3 7 1150 1169 -568 1136 792 2 <2e-16 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 122.3.3 Cross-Level Interaction
Helps for “failed to converge”: https://cran.r-project.org/web/packages/lme4/vignettes/lmerperf.html
fit_glmer_4 <- lme4::glmer(cbind(res, denom - res) ~ respisrr + mode +
saliency + yr77 +
(1|source) + (0 + tel|source) + (0+ mail|source),
data = df_long,
family = binomial,
control=glmerControl(optimizer="bobyqa",
optCtrl=list(maxfun=100000)))Generalized linear mixed model fit by maximum likelihood (Laplace
Approximation) [glmerMod]
Family: binomial ( logit )
Formula: cbind(res, denom - res) ~ respisrr + mode + saliency + yr77 +
(1 | source) + (0 + tel | source) + (0 + mail | source)
Data: df_long
Control: glmerControl(optimizer = "bobyqa", optCtrl = list(maxfun = 1e+05))
AIC BIC logLik -2*log(L) df.resid
1138 1164 -559 1118 95
Scaled residuals:
Min 1Q Median 3Q Max
-1.2213 -0.1306 0.0093 0.2001 1.0561
Random effects:
Groups Name Variance Std.Dev.
source (Intercept) 0.516 0.719
source.1 tel 0.252 0.502
source.2 mail 0.409 0.640
Number of obs: 105, groups: source, 48
Fixed effects:
Estimate Std. Error z value Pr(>|z|)
(Intercept) 0.3438 0.2841 1.21 0.22622
respisrrRr 0.2527 0.2130 1.19 0.23531
modeTel -0.1558 0.0940 -1.66 0.09740 .
modeMail -0.5037 0.1392 -3.62 0.00029 ***
saliencySome 0.6800 0.2963 2.30 0.02172 *
saliencyHigh 1.3583 0.3320 4.09 4.3e-05 ***
yr77 -0.0230 0.0118 -1.94 0.05249 .
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Correlation of Fixed Effects:
(Intr) rspsrR modeTl modeMl slncyS slncyH
respisrrRr -0.437
modeTel -0.121 0.085
modeMail -0.115 0.102 0.129
saliencySom -0.722 -0.111 0.035 -0.003
saliencyHgh -0.689 0.016 0.058 0.028 0.644
yr77 0.136 -0.192 0.002 0.054 -0.078 0.051fit_glmer_5 <- lme4::glmer(cbind(res, denom - res) ~ respisrr + mode +
saliency + mode*yr77 +
(1|source) + (0 + tel|source) + (0+ mail|source),
data = df_long,
family = binomial,
control=glmerControl(optimizer="bobyqa",
optCtrl=list(maxfun=100000)))Generalized linear mixed model fit by maximum likelihood (Laplace
Approximation) [glmerMod]
Family: binomial ( logit )
Formula: cbind(res, denom - res) ~ respisrr + mode + saliency + mode *
yr77 + (1 | source) + (0 + tel | source) + (0 + mail | source)
Data: df_long
Control: glmerControl(optimizer = "bobyqa", optCtrl = list(maxfun = 1e+05))
AIC BIC logLik -2*log(L) df.resid
1131 1163 -554 1107 93
Scaled residuals:
Min 1Q Median 3Q Max
-0.8668 -0.1418 0.0197 0.1920 1.0444
Random effects:
Groups Name Variance Std.Dev.
source (Intercept) 0.541 0.736
source.1 tel 0.224 0.473
source.2 mail 0.321 0.567
Number of obs: 105, groups: source, 48
Fixed effects:
Estimate Std. Error z value Pr(>|z|)
(Intercept) 0.3505 0.2882 1.22 0.2239
respisrrRr 0.2629 0.2105 1.25 0.2117
modeTel -0.1797 0.0909 -1.98 0.0479 *
modeMail -0.4940 0.1263 -3.91 9.2e-05 ***
saliencySome 0.6876 0.3013 2.28 0.0225 *
saliencyHigh 1.3784 0.3389 4.07 4.8e-05 ***
yr77 -0.0287 0.0122 -2.34 0.0192 *
modeTel:yr77 0.0209 0.0101 2.06 0.0389 *
modeMail:yr77 0.0363 0.0134 2.70 0.0069 **
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Correlation of Fixed Effects:
(Intr) rspsrR modeTl modeMl slncyS slncyH yr77 mdT:77
respisrrRr -0.429
modeTel -0.128 0.089
modeMail -0.115 0.098 0.145
saliencySom -0.730 -0.101 0.039 0.002
saliencyHgh -0.692 0.019 0.064 0.028 0.646
yr77 0.135 -0.193 0.006 0.050 -0.080 0.041
modeTl:yr77 0.037 -0.011 -0.143 -0.026 -0.024 -0.026 -0.052
modeMl:yr77 0.018 -0.043 -0.015 0.020 0.038 0.030 -0.160 0.04622.3.3.1 Compare
| Main Effect | Intraction | ||||
|---|---|---|---|---|---|---|
Variable | b | (SE) | p | b | (SE) | p |
(Intercept) | 0.34 | (0.28) | .226 | 0.35 | (0.29) | .224 |
respisrr | ||||||
Cr | — | — | — | — | ||
Rr | 0.25 | (0.21) | .235 | 0.26 | (0.21) | .212 |
mode | ||||||
Ftf | — | — | — | — | ||
Tel | -0.16 | (0.09) | .097 | -0.18 | (0.09) | .048* |
-0.50 | (0.14) | < .001*** | -0.49 | (0.13) | < .001*** | |
saliency | ||||||
Not | — | — | — | — | ||
Some | 0.68 | (0.30) | .022* | 0.69 | (0.30) | .022* |
High | 1.36 | (0.33) | < .001*** | 1.38 | (0.34) | < .001*** |
yr77 | -0.02 | (0.01) | .052 | -0.03 | (0.01) | .019* |
mode * yr77 | ||||||
Tel * yr77 | 0.02 | (0.01) | .039* | |||
Mail * yr77 | 0.04 | (0.01) | .007** | |||
source.var__(Intercept) | 0.52 | 0.54 | ||||
source.1.var__tel | 0.25 | 0.22 | ||||
source.2.var__mail | 0.41 | 0.32 | ||||
AIC | 1,138 | 1,131 | ||||
BIC | 1,164 | 1,163 | ||||
Log-likelihood | -559 | -554 | ||||
Note. | ||||||
* p < .05. ** p < .01. *** p < .001. | ||||||
Model | AIC | BIC | RMSE |
| |
|---|---|---|---|---|---|
Conditional | Marginal | ||||
Main Effect | 1137.61 | 1164.15 | 0.01 | .996 | .362 |
Intraction | 1131.18 | 1163.03 | 0.01 | .996 | .350 |
Note. Larger values indicated better performance. Smaller values indicated better performance for Akaike's Information Criteria (AIC), Bayesian information criteria (BIC), and Root Mean Squared Error (RMSE). | |||||
Data: df_long
Models:
fit_glmer_4: cbind(res, denom - res) ~ respisrr + mode + saliency + yr77 + (1 | source) + (0 + tel | source) + (0 + mail | source)
fit_glmer_5: cbind(res, denom - res) ~ respisrr + mode + saliency + mode * yr77 + (1 | source) + (0 + tel | source) + (0 + mail | source)
npar AIC BIC logLik -2*log(L) Chisq Df Pr(>Chisq)
fit_glmer_4 10 1138 1164 -559 1118
fit_glmer_5 12 1131 1163 -554 1107 10.4 2 0.0054 **
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 122.3.4 Predicted Probabilities
22.3.4.1 At the Mean Year (1977)
fit_glmer_5 %>%
emmeans::emmeans(~ mode*saliency,
at = list(yr77 = 0,
saliency = c("Not", "High")),
type = "response") mode saliency prob SE df asymp.LCL asymp.UCL
Ftf Not 0.618 0.0616 Inf 0.493 0.730
Tel Not 0.575 0.0653 Inf 0.445 0.695
Mail Not 0.497 0.0699 Inf 0.363 0.631
Ftf High 0.865 0.0258 Inf 0.806 0.908
Tel High 0.843 0.0314 Inf 0.771 0.895
Mail High 0.797 0.0402 Inf 0.707 0.864
Results are averaged over the levels of: respisrr
Confidence level used: 0.95
Intervals are back-transformed from the logit scale 22.3.4.2 Each Mode Across Years
fit_glmer_5 %>%
emmeans::emmeans(~ mode*yr77,
at = list(yr77 = c(1950-1977,
1977-1977,
1990-1977)),
type = "response") mode yr77 prob SE df asymp.LCL asymp.UCL
Ftf -27 0.875 0.0372 Inf 0.782 0.932
Tel -27 0.769 0.0777 Inf 0.585 0.887
Mail -27 0.616 0.1080 Inf 0.396 0.797
Ftf 0 0.763 0.0232 Inf 0.715 0.806
Tel 0 0.729 0.0291 Inf 0.669 0.782
Mail 0 0.663 0.0371 Inf 0.587 0.731
Ftf 13 0.689 0.0463 Inf 0.592 0.772
Tel 13 0.709 0.0526 Inf 0.596 0.801
Mail 13 0.685 0.0629 Inf 0.551 0.794
Results are averaged over the levels of: respisrr, saliency
Confidence level used: 0.95
Intervals are back-transformed from the logit scale fit_glmer_5 %>%
emmeans::emmeans(~ mode*yr77,
at = list(yr77 = c(1950-1977,
1977-1977,
1990-1977)),
type = "response") %>%
data.frame() %>%
dplyr::mutate(year = as.character(1977 + yr77)) %>%
dplyr::mutate(mode = mode %>%
forcats::fct_recode("Face-to-Face" = "Ftf",
"Telephone" = "Tel",
"Mail" = "Mail")) %>%
dplyr::mutate(across(c(prob, asymp.LCL, asymp.UCL),
~apaSupp::p_num(.x, stars = FALSE))) %>%
dplyr::mutate(cell = glue::glue("{prob} [{asymp.LCL}, {asymp.UCL}]")) %>%
dplyr::select(mode, year, cell) %>%
tidyr::pivot_wider(names_from = year,
values_from = cell) %>%
flextable::flextable() %>%
apaSupp::theme_apa(caption = "Predicted Response Rates for Each Mode at Key Years, [95% CI]",
general_note = "Predications are for 'some' saliency.")mode | 1950 | 1977 | 1990 |
|---|---|---|---|
Face-to-Face | .875 [ .782 , .932 ] | .763 [ .715 , .806 ] | .689 [ .592 , .772 ] |
Telephone | .769 [ .585 , .887 ] | .729 [ .669 , .782 ] | .709 [ .596 , .801 ] |
.616 [ .396 , .797 ] | .663 [ .587 , .731 ] | .685 [ .551 , .794 ] | |
Note. Predications are for 'some' saliency. | |||
22.3.4.3 Pairwise Comparisons
fit_glmer_5 %>%
emmeans::emmeans(~ mode*saliency|yr77,
at = list(saliency = "Some",
yr77 = c(1950-1977,
1977-1977,
1990-1977)),
type = "response") %>%
pairs(adjust = "none")yr77 = -27:
contrast odds.ratio SE df null z.ratio p.value
Ftf Some / Tel Some 2.11 0.632 Inf 1 2.480 0.0130
Ftf Some / Mail Some 4.36 1.660 Inf 1 3.860 <.0001
Tel Some / Mail Some 2.07 0.973 Inf 1 1.550 0.1210
yr77 = 0:
contrast odds.ratio SE df null z.ratio p.value
Ftf Some / Tel Some 1.20 0.109 Inf 1 1.980 0.0480
Ftf Some / Mail Some 1.64 0.207 Inf 1 3.910 <.0001
Tel Some / Mail Some 1.37 0.198 Inf 1 2.170 0.0300
yr77 = 13:
contrast odds.ratio SE df null z.ratio p.value
Ftf Some / Tel Some 0.91 0.136 Inf 1 -0.620 0.5360
Ftf Some / Mail Some 1.02 0.222 Inf 1 0.100 0.9170
Tel Some / Mail Some 1.12 0.287 Inf 1 0.450 0.6530
Results are averaged over the levels of: respisrr
Tests are performed on the log odds ratio scale fit_glmer_5 %>%
emmeans::emmeans(~ mode*saliency|yr77,
at = list(saliency = "Some",
yr77 = c(1950-1977,
1977-1977,
1990-1977)),
type = "response") %>%
pairs(adjust = "none") %>%
data.frame() %>%
dplyr::mutate(yr77 = as.numeric(as.character(yr77))) %>%
dplyr::mutate(year = as.character(1977 + yr77)) %>%
dplyr::mutate(contrast = contrast %>%
forcats::fct_recode("Face-to-Face vs. Telephone" = "Ftf Some / Tel Some",
"Face-to-Face vs. Mail" = "Ftf Some / Mail Some",
"Telephone vs. Mail" = "Tel Some / Mail Some")) %>%
dplyr::mutate(p.adj = p.adjust(p.value, method = "fdr")) %>%
dplyr::mutate(across(c(p.value, p.adj), apaSupp::p_num)) %>%
dplyr::select(Year = year,
"Comparing Modes" = contrast,
OR = odds.ratio,
p_Unadj = p.value,
p_Adj = p.adj) %>%
flextable::as_grouped_data(groups = "Year") %>%
flextable::flextable() %>%
flextable::separate_header() %>%
apaSupp::theme_apa(caption = "Pairwise Comparisons Between Survey Mode at Key Years",
general_note = "Significance is given both unadjusted (Unadj) and adjusted (Adj) via the method of Benjamini, Hochberg, and Yekutieli to control the false discovery rate (FDR).")Year | Comparing Modes | OR | p | |
|---|---|---|---|---|
Unadj | Adj | |||
1950 | ||||
Face-to-Face vs. Telephone | 2.11 | .013* | .040* | |
Face-to-Face vs. Mail | 4.36 | < .001*** | < .001*** | |
Telephone vs. Mail | 2.07 | .121 | .181 | |
1977 | ||||
Face-to-Face vs. Telephone | 1.20 | .048* | .086 | |
Face-to-Face vs. Mail | 1.64 | < .001*** | < .001*** | |
Telephone vs. Mail | 1.37 | .030* | .067 | |
1990 | ||||
Face-to-Face vs. Telephone | 0.91 | .536 | .689 | |
Face-to-Face vs. Mail | 1.02 | .917 | .917 | |
Telephone vs. Mail | 1.12 | .653 | .735 | |
Note. Significance is given both unadjusted (Unadj) and adjusted (Adj) via the method of Benjamini, Hochberg, and Yekutieli to control the false discovery rate (FDR). | ||||
22.3.5 Plot
22.3.5.1 Simple
interactions::interact_plot(model = fit_glmer_5,
pred = yr77,
modx = mode,
interval = TRUE,
int.width = .685)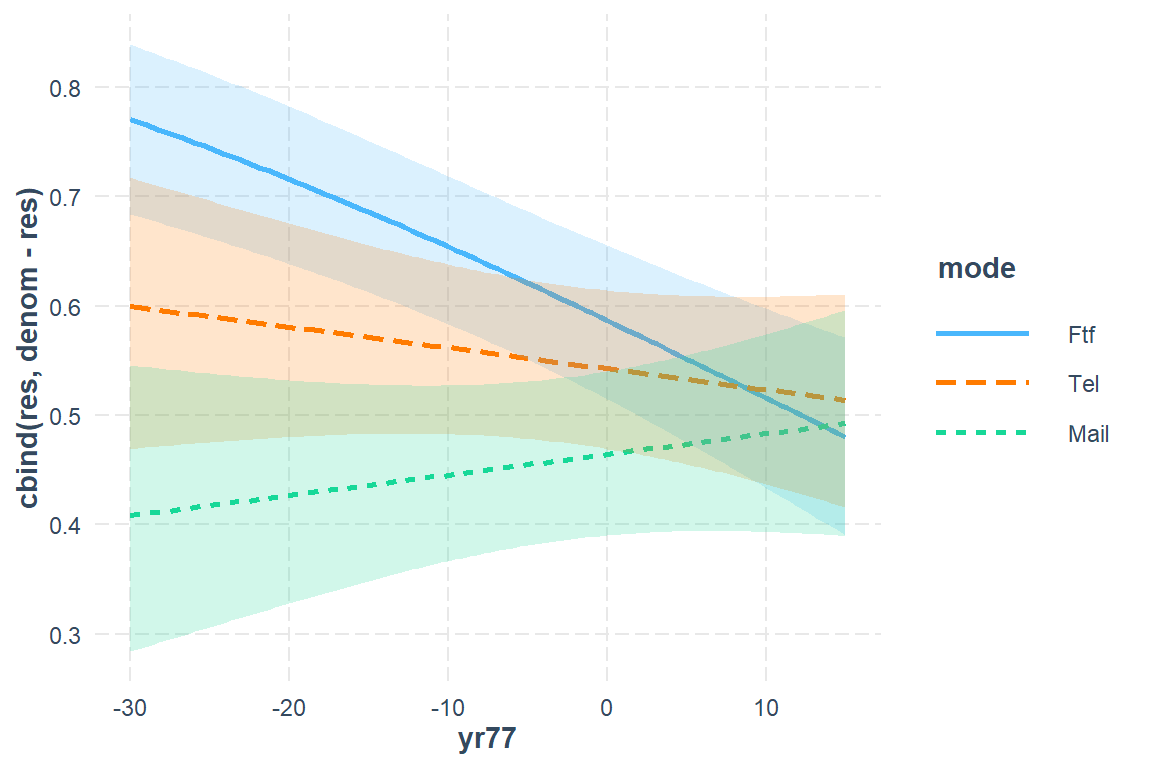
22.3.5.2 Publishable
fit_glmer_5 %>%
emmeans::emmeans(~ yr77*mode,
at = list(yr77 = seq(from = 1947 - 1977,
to = 1992 - 1977,
by = 1)),
type = "response",
level = .685) %>%
data.frame() %>%
dplyr::mutate(year = 1977 + yr77) %>%
dplyr::mutate(mode = mode %>%
forcats::fct_recode("Face-to-Face" = "Ftf",
"Telephone" = "Tel",
"Mail" = "Mail")) %>%
ggplot(aes(x = year,
y = prob,
linetype = mode)) +
# geom_ribbon(aes(ymin = asymp.LCL,
# ymax = asymp.UCL),
# alpha= .15) +
geom_line(linewidth = 1) +
labs(x = NULL,
y = "Predicted Response Rate",
linetype = "Collection Mode") +
theme_bw() +
geom_vline(xintercept = c(1950, 1977, 1990),
linewidth = 3,
alpha = .2) +
scale_linetype_manual(values = c("solid", "longdash", "dotted")) +
theme(legend.position = "inside",
legend.position.inside = c(.92, 1.0),
legend.justification = c(1.1, 1.1),
legend.background = element_rect(color = "black"),
legend.key.width = unit(1.5, "cm"))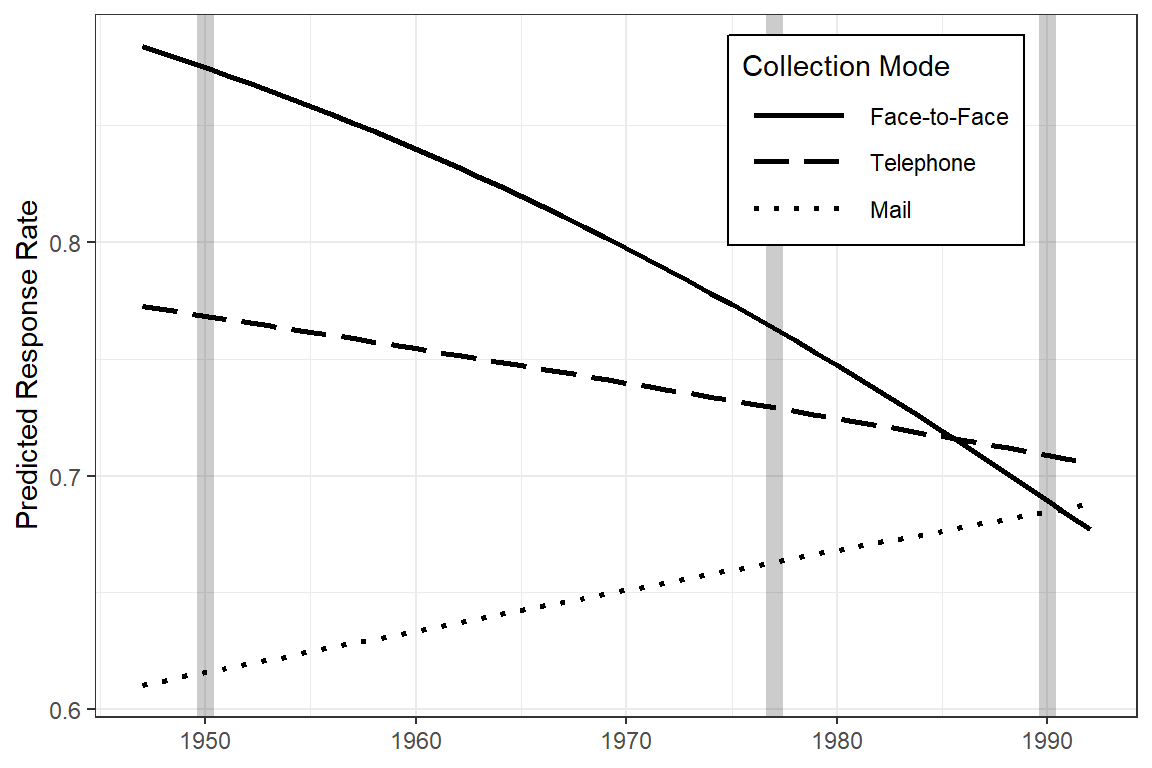
Figure 22.2
Predicted Response Rates Over the Years (Hox figure 6.2 on page 122)