16 GEE, Binary Outcome: Respiratory Illness
16.1 PREPARATION
16.1.1 Load Packages
install.packages("devtools")
devtools::install_github("sarbearschwartz/apaSupp") # updated 10/14/2025Load/activate these packages
16.1.2 Background
This dataset was used as an example in Chapter 11 of “A Handbook of Statistical Analysis using R” by Brian S. Everitt and Torsten Hothorn. The authors include this data set in their HSAUR package on
CRAN.
The Background
In each of two centers, eligible patients were randomly assigned to active treatment or placebo. During the treatment, the respiratory status (categorized poor or good) was determined at each of four, monthly visits. The trial recruited 111 participants (54 in the active group, 57 in the placebo group) and there were no missing data for either the responses or the covariates.
The Research Question
The question of interest is to assess whether the treatment is effective and to estimate its effect.
Note: We are NOT interested in change over time, but rather mean differences in the treatment group compared to the placebo group, net of any potential confounding due to age, sex, and site.
The Data
Note that the data (555 observations on the following 7 variables) are in long form, i.e, repeated measurements are stored as additional rows in the data frame.
Indicators
subjectthe patient ID, a factor with levels 1 to 111centrethe study center, a factor with levels 1 and 2
monththe month, each patient was examined at months 0, 1, 2, 3 and 4Outcome or dependent variable
statusthe respiratory status (response variable), a factor with levels poor and goodMain predictor or independent variable of interest
treatmentthe treatment arm, a factor with levels placebo and treatmentTime-invariant Covariates to control for
sexa factor with levels female and male
agethe age of the patient
16.1.3 Read in the data
'data.frame': 555 obs. of 7 variables:
$ centre : Factor w/ 2 levels "1","2": 1 1 1 1 1 1 1 1 1 1 ...
$ treatment: Factor w/ 2 levels "placebo","treatment": 1 1 1 1 1 1 1 1 1 1 ...
$ sex : Factor w/ 2 levels "female","male": 1 1 1 1 1 1 1 1 1 1 ...
$ age : num 46 46 46 46 46 28 28 28 28 28 ...
$ status : Factor w/ 2 levels "poor","good": 1 1 1 1 1 1 1 1 1 1 ...
$ month : Ord.factor w/ 5 levels "0"<"1"<"2"<"3"<..: 1 2 3 4 5 1 2 3 4 5 ...
$ subject : Factor w/ 111 levels "1","2","3","4",..: 1 1 1 1 1 2 2 2 2 2 ...Rows: 555
Columns: 7
$ centre <fct> 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, …
$ treatment <fct> placebo, placebo, placebo, placebo, placebo, placebo, placeb…
$ sex <fct> female, female, female, female, female, female, female, fema…
$ age <dbl> 46, 46, 46, 46, 46, 28, 28, 28, 28, 28, 23, 23, 23, 23, 23, …
$ status <fct> poor, poor, poor, poor, poor, poor, poor, poor, poor, poor, …
$ month <ord> 0, 1, 2, 3, 4, 0, 1, 2, 3, 4, 0, 1, 2, 3, 4, 0, 1, 2, 3, 4, …
$ subject <fct> 1, 1, 1, 1, 1, 2, 2, 2, 2, 2, 3, 3, 3, 3, 3, 4, 4, 4, 4, 4, …16.1.4 Wide Format
Wide format has one line per participant.
df_resp_wide <- respiratory %>%
tidyr::spread(key = month,
value = status,
sep = "_") %>%
dplyr::rename("BL_status" = "month_0") %>%
dplyr::arrange(subject) %>%
dplyr::select(subject, centre,
sex, age, treatment,
BL_status, starts_with("month")) %>%
dplyr::mutate_if(is.factor, ~forcats::fct_relabel(.x, stringr::str_to_title))Rows: 111
Columns: 10
$ subject <fct> 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 1…
$ centre <fct> 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, …
$ sex <fct> Female, Female, Female, Female, Male, Female, Female, Female…
$ age <dbl> 46, 28, 23, 44, 13, 34, 43, 28, 31, 37, 30, 14, 23, 30, 20, …
$ treatment <fct> Placebo, Placebo, Treatment, Placebo, Placebo, Treatment, Pl…
$ BL_status <fct> Poor, Poor, Good, Good, Good, Poor, Poor, Poor, Good, Good, …
$ month_1 <fct> Poor, Poor, Good, Good, Good, Poor, Good, Poor, Good, Poor, …
$ month_2 <fct> Poor, Poor, Good, Good, Good, Poor, Poor, Poor, Good, Good, …
$ month_3 <fct> Poor, Poor, Good, Good, Good, Poor, Good, Poor, Good, Good, …
$ month_4 <fct> Poor, Poor, Good, Poor, Good, Poor, Good, Poor, Good, Poor, …df_resp_wide %>%
psych::headTail(top = 10, bottom = 10) %>%
flextable::flextable() %>%
apaSupp::theme_apa(caption = "Wide Format")subject | centre | sex | age | treatment | BL_status | month_1 | month_2 | month_3 | month_4 |
|---|---|---|---|---|---|---|---|---|---|
1 | 1 | Female | 46 | Placebo | Poor | Poor | Poor | Poor | Poor |
2 | 1 | Female | 28 | Placebo | Poor | Poor | Poor | Poor | Poor |
3 | 1 | Female | 23 | Treatment | Good | Good | Good | Good | Good |
4 | 1 | Female | 44 | Placebo | Good | Good | Good | Good | Poor |
5 | 1 | Male | 13 | Placebo | Good | Good | Good | Good | Good |
6 | 1 | Female | 34 | Treatment | Poor | Poor | Poor | Poor | Poor |
7 | 1 | Female | 43 | Placebo | Poor | Good | Poor | Good | Good |
8 | 1 | Female | 28 | Treatment | Poor | Poor | Poor | Poor | Poor |
9 | 1 | Female | 31 | Treatment | Good | Good | Good | Good | Good |
10 | 1 | Female | 37 | Placebo | Good | Poor | Good | Good | Poor |
... | |||||||||
102 | 2 | Female | 48 | Placebo | Good | Good | Poor | Poor | Poor |
103 | 2 | Female | 20 | Treatment | Poor | Good | Good | Good | Good |
104 | 2 | Male | 39 | Placebo | Good | Poor | Good | Poor | Poor |
105 | 2 | Female | 28 | Treatment | Poor | Good | Poor | Poor | Poor |
106 | 2 | Male | 38 | Placebo | Poor | Poor | Poor | Poor | Poor |
107 | 2 | Female | 43 | Treatment | Good | Good | Good | Good | Good |
108 | 2 | Male | 39 | Treatment | Poor | Good | Good | Good | Good |
109 | 2 | Female | 68 | Treatment | Poor | Good | Good | Good | Good |
110 | 2 | Male | 63 | Treatment | Good | Good | Good | Good | Good |
111 | 2 | Female | 31 | Treatment | Good | Good | Good | Good | Good |
16.1.5 Long Format
Long format has one line per observation.
df_resp_long <- df_resp_wide%>%
tidyr::gather(key = month,
value = status,
starts_with("month")) %>%
dplyr::mutate(month = str_sub(month, start = -1) %>% as.numeric) %>%
dplyr::mutate(status = case_when(status == "Poor" ~ 0,
status == "Good" ~ 1)) %>%
dplyr::arrange(subject, month) %>%
dplyr::select(subject, centre, sex, age, treatment, BL_status, month, status)Rows: 444
Columns: 8
$ subject <fct> 1, 1, 1, 1, 2, 2, 2, 2, 3, 3, 3, 3, 4, 4, 4, 4, 5, 5, 5, 5, …
$ centre <fct> 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, …
$ sex <fct> Female, Female, Female, Female, Female, Female, Female, Fema…
$ age <dbl> 46, 46, 46, 46, 28, 28, 28, 28, 23, 23, 23, 23, 44, 44, 44, …
$ treatment <fct> Placebo, Placebo, Placebo, Placebo, Placebo, Placebo, Placeb…
$ BL_status <fct> Poor, Poor, Poor, Poor, Poor, Poor, Poor, Poor, Good, Good, …
$ month <dbl> 1, 2, 3, 4, 1, 2, 3, 4, 1, 2, 3, 4, 1, 2, 3, 4, 1, 2, 3, 4, …
$ status <dbl> 0, 0, 0, 0, 0, 0, 0, 0, 1, 1, 1, 1, 1, 1, 1, 0, 1, 1, 1, 1, …df_resp_long %>%
psych::headTail(top = 10, bottom = 10) %>%
flextable::flextable() %>%
apaSupp::theme_apa(caption = "Long Format")subject | centre | sex | age | treatment | BL_status | month | status |
|---|---|---|---|---|---|---|---|
1 | 1 | Female | 46 | Placebo | Poor | 1 | 0 |
1 | 1 | Female | 46 | Placebo | Poor | 2 | 0 |
1 | 1 | Female | 46 | Placebo | Poor | 3 | 0 |
1 | 1 | Female | 46 | Placebo | Poor | 4 | 0 |
2 | 1 | Female | 28 | Placebo | Poor | 1 | 0 |
2 | 1 | Female | 28 | Placebo | Poor | 2 | 0 |
2 | 1 | Female | 28 | Placebo | Poor | 3 | 0 |
2 | 1 | Female | 28 | Placebo | Poor | 4 | 0 |
3 | 1 | Female | 23 | Treatment | Good | 1 | 1 |
3 | 1 | Female | 23 | Treatment | Good | 2 | 1 |
... | ... | ... | |||||
109 | 2 | Female | 68 | Treatment | Poor | 3 | 1 |
109 | 2 | Female | 68 | Treatment | Poor | 4 | 1 |
110 | 2 | Male | 63 | Treatment | Good | 1 | 1 |
110 | 2 | Male | 63 | Treatment | Good | 2 | 1 |
110 | 2 | Male | 63 | Treatment | Good | 3 | 1 |
110 | 2 | Male | 63 | Treatment | Good | 4 | 1 |
111 | 2 | Female | 31 | Treatment | Good | 1 | 1 |
111 | 2 | Female | 31 | Treatment | Good | 2 | 1 |
111 | 2 | Female | 31 | Treatment | Good | 3 | 1 |
111 | 2 | Female | 31 | Treatment | Good | 4 | 1 |
16.2 EXPLORATORY DATA ANLAYSIS
16.2.1 Summary Statistics
16.2.1.1 Demographics and Baseline Measure
Notice that numerical summaries are computed for all variables, even the categorical variables (i.e. factors). The have an * after the variable name to remind you that the mean, sd, and se are of limited use.
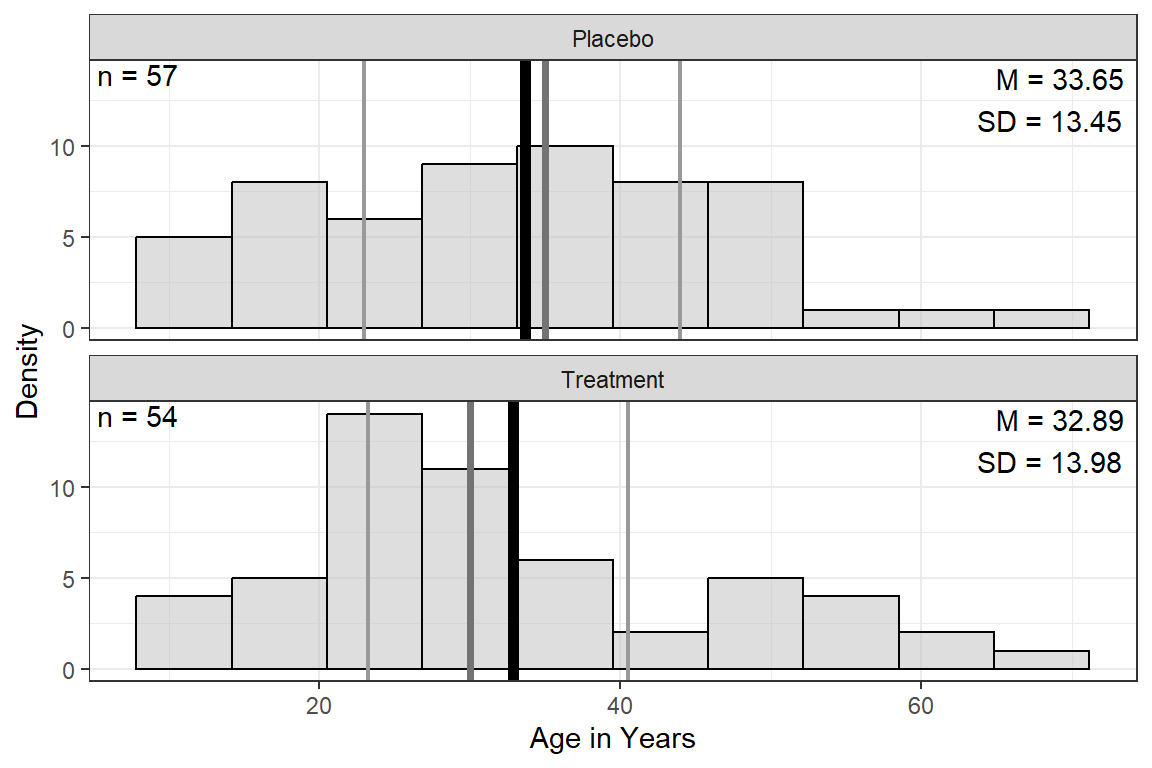
Notice: the mean age is 33
NA | M | SD | min | Q1 | Mdn | Q3 | max | |
|---|---|---|---|---|---|---|---|---|
age | 0 | 33.28 | 13.65 | 11.00 | 23.00 | 31.00 | 43.00 | 68.00 |
Note. N = 111. NA = not available or missing; Mdn = median; Q1 = 25th percentile; Q3 = 75th percentile. | ||||||||
df_resp_wide %>%
dplyr::select(treatment,
"Center" = centre,
"Sex" = sex,
"Age" = age,
"Baseline Status" = BL_status) %>%
apaSupp::tab1(split = "treatment",
caption = "Participant Demographics",
p_note = "apa1")
| Total | Placebo | Treatment | p-value |
|---|---|---|---|---|
Center | 1.000 | |||
1 | 56 (50.5%) | 29 (50.9%) | 27 (50.0%) | |
2 | 55 (49.5%) | 28 (49.1%) | 27 (50.0%) | |
Sex | .028* | |||
Female | 88 (79.3%) | 40 (70.2%) | 48 (88.9%) | |
Male | 23 (20.7%) | 17 (29.8%) | 6 (11.1%) | |
Age | 33.28 (13.65) | 33.65 (13.45) | 32.89 (13.98) | .771 |
Baseline Status | 1.000 | |||
Poor | 61 (55.0%) | 31 (54.4%) | 30 (55.6%) | |
Good | 50 (45.0%) | 26 (45.6%) | 24 (44.4%) | |
Note. Continuous variables are summarized with means (SD) and significant group differences assessed via independent t-tests. Categorical variables are summarized with counts (%) and significant group differences assessed via Chi-squared tests for independence. | ||||
* p < .05. | ||||
16.2.1.2 Status Over Time
df_resp_wide %>%
dplyr::select(treatment,
"Month One" = month_1,
"Month Two" = month_2,
"Month Three" = month_3,
"Month Four" = month_4) %>%
apaSupp::tab1(split = "treatment",
caption = "Respiratory Status Over Time",
p_note = "apa2")
| Total | Placebo | Treatment | p-value |
|---|---|---|---|---|
Month One | .060 | |||
Poor | 46 (41.4%) | 29 (50.9%) | 17 (31.5%) | |
Good | 65 (58.6%) | 28 (49.1%) | 37 (68.5%) | |
Month Two | .002** | |||
Poor | 51 (45.9%) | 35 (61.4%) | 16 (29.6%) | |
Good | 60 (54.1%) | 22 (38.6%) | 38 (70.4%) | |
Month Three | .008** | |||
Poor | 46 (41.4%) | 31 (54.4%) | 15 (27.8%) | |
Good | 65 (58.6%) | 26 (45.6%) | 39 (72.2%) | |
Month Four | .068 | |||
Poor | 52 (46.8%) | 32 (56.1%) | 20 (37.0%) | |
Good | 59 (53.2%) | 25 (43.9%) | 34 (63.0%) | |
Note. . . | ||||
** p < .01. | ||||
Correlation between repeated observations:
df_resp_wide %>%
dplyr::select(starts_with("month")) %>%
dplyr::mutate_all(function(x) x == "Good") %>%
cor() %>%
corrplot::corrplot.mixed()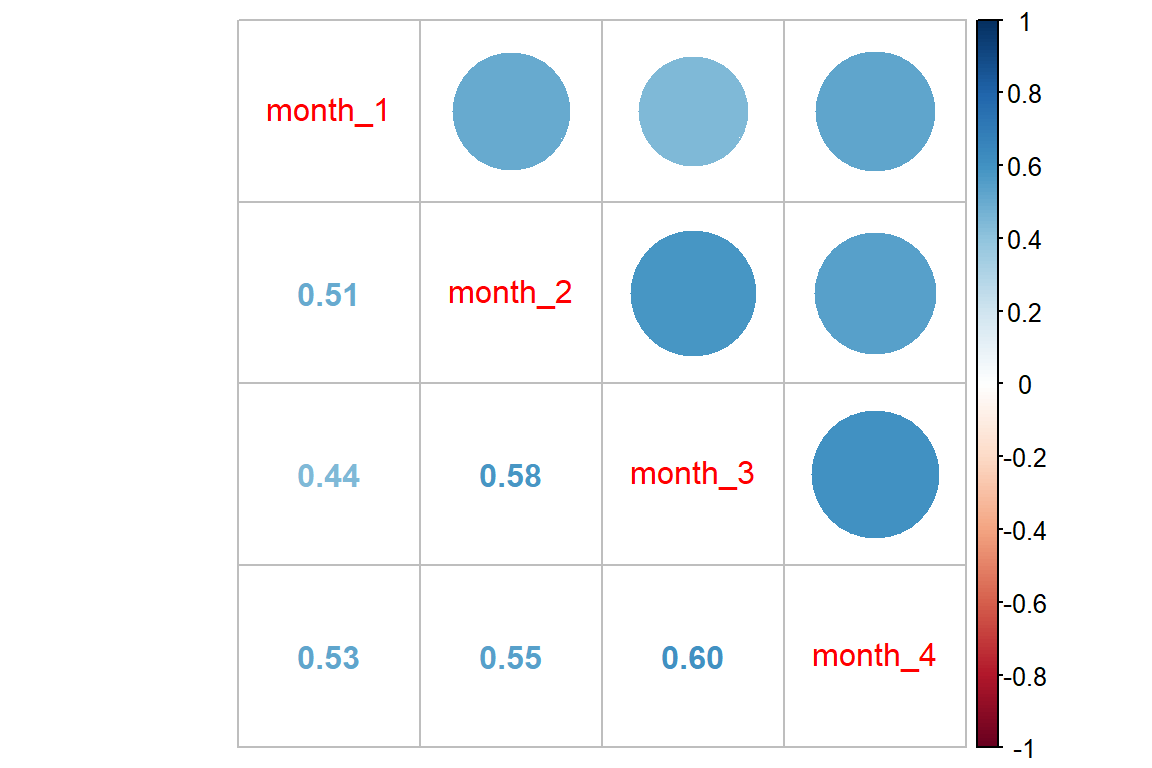
16.2.2 Visualization
16.2.2.2 Status by Age
df_resp_wide %>%
dplyr::mutate(n_good = furniture::rowsums(month_1 == "Good",
month_2 == "Good",
month_3 == "Good",
month_4 == "Good")) %>%
ggplot(aes(x = age,
y = n_good)) +
geom_count() +
geom_smooth() +
theme_bw() +
labs(x = "Age in Years",
y = "Number of Visits out of Four\nwith 'Good' Respiration")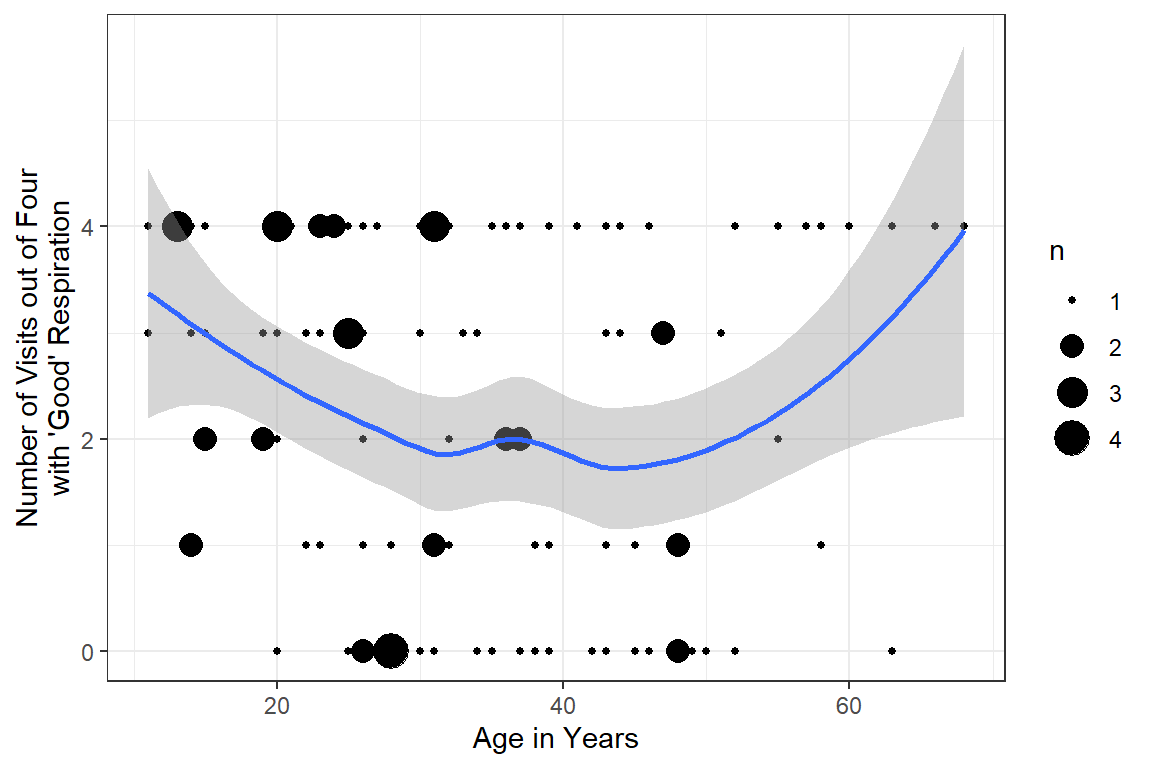
16.2.2.3 Status Over Time
It appears that status is fairly constant over time.
df_resp_long %>%
dplyr::group_by(treatment, month) %>%
dplyr::summarise(n = n(),
prop_good = mean(status),
prop_sd = sd(status),
prop_se = prop_sd/sqrt(n)) %>%
ggplot(aes(x = month,
y = prop_good,
group = treatment,
color = treatment)) +
geom_errorbar(aes(ymin = prop_good - prop_se,
ymax = prop_good + prop_se),
width = .25,
show.legend = FALSE) +
geom_point(aes(shape = treatment),
size = 2) +
geom_line(aes(linetype = treatment)) +
theme_bw() +
labs(x = "Time, months",
y = "Observed Proportion of Participants\nwith 'Good' Respiratory Status",
color = NULL,
shape = NULL,
linetype = NULL) +
scale_linetype_manual(values = c("solid", "longdash")) +
theme(legend.position = "bottom",
legend.key.width = unit(1.5, "cm"))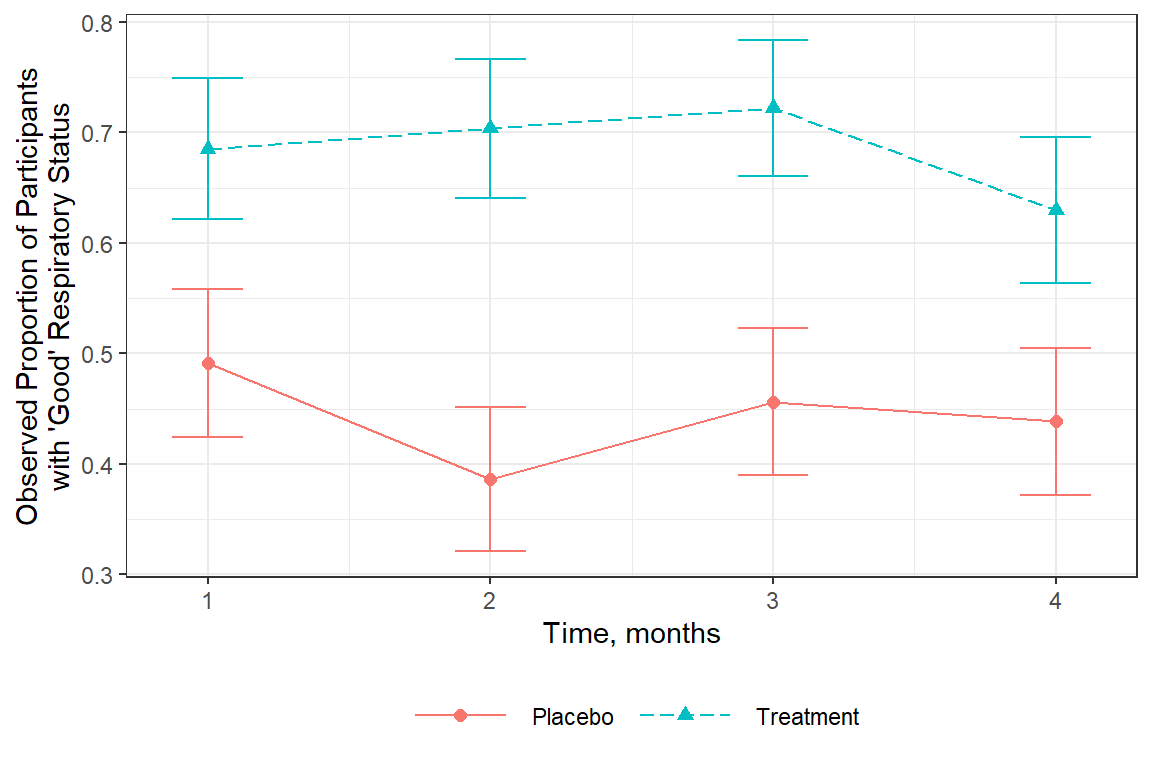
It is NOT the purpose of this analysis to investigate change over time!
Since status is largely stable over time, no linear (or even
quadratic) effect of the month variable will be
modeled.
Instead, the four observations on each subject are treated as correlated (at least with non-independent correlation structure in GEE), but no time trend will be included.
16.3 Generalized Estimating Equations (GEE)
Again, since participants age is likely to be a risk at either end of the spectrum, the potential quadratic effect of age will be modeled. Age is also being grand-mean centered to make the intercept more meaningful.
16.3.1 Indepdendence
Using an "independence" correlation structure is equivalent to using a GLM analysis (logistic regression in this case) and is never appropriate for repeated measures data. It is only being done here for comparison purposes.
fit_geeglm_in <- geepack::geeglm(status ~ centre + treatment + sex + BL_status +
I(age-33) + I((age-33)^2),
data = df_resp_long,
family = binomial(link = "logit"),
id = subject,
waves = month,
corstr = "independence",
scale.fix = TRUE)
summary(fit_geeglm_in)
Call:
geepack::geeglm(formula = status ~ centre + treatment + sex +
BL_status + I(age - 33) + I((age - 33)^2), family = binomial(link = "logit"),
data = df_resp_long, id = subject, waves = month, corstr = "independence",
scale.fix = TRUE)
Coefficients:
Estimate Std.err Wald Pr(>|W|)
(Intercept) -1.9685725 0.4457014 19.508 1.00e-05 ***
centre2 0.5347938 0.3795760 1.985 0.158858
treatmentTreatment 1.3561814 0.3777999 12.886 0.000331 ***
sexMale 0.4263433 0.4832337 0.778 0.377630
BL_statusGood 1.9193401 0.3772812 25.881 3.63e-07 ***
I(age - 33) -0.0368535 0.0150120 6.027 0.014091 *
I((age - 33)^2) 0.0025169 0.0007592 10.989 0.000917 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Correlation structure = independence
Scale is fixed.
Number of clusters: 111 Maximum cluster size: 4 The results for GEE fit with the independence correlation structure produces results that are nearly identical to the GLM model.
16.3.2 Exchangeable
fit_geeglm_ex <- geepack::geeglm(status ~ centre + treatment + sex + BL_status +
I(age-33) + I((age-33)^2),
data = df_resp_long,
family = binomial(link = "logit"),
id = subject,
waves = month,
corstr = "exchangeable",
scale.fix = TRUE)
summary(fit_geeglm_ex)
Call:
geepack::geeglm(formula = status ~ centre + treatment + sex +
BL_status + I(age - 33) + I((age - 33)^2), family = binomial(link = "logit"),
data = df_resp_long, id = subject, waves = month, corstr = "exchangeable",
scale.fix = TRUE)
Coefficients:
Estimate Std.err Wald Pr(>|W|)
(Intercept) -1.968572 0.445701 19.51 1.0e-05 ***
centre2 0.534794 0.379576 1.99 0.15886
treatmentTreatment 1.356181 0.377800 12.89 0.00033 ***
sexMale 0.426343 0.483234 0.78 0.37763
BL_statusGood 1.919340 0.377281 25.88 3.6e-07 ***
I(age - 33) -0.036854 0.015012 6.03 0.01409 *
I((age - 33)^2) 0.002517 0.000759 10.99 0.00092 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Correlation structure = exchangeable
Scale is fixed.
Link = identity
Estimated Correlation Parameters:
Estimate Std.err
alpha 0.312 0.0521
Number of clusters: 111 Maximum cluster size: 4 16.3.3 Auto-Regressive
fit_geeglm_ar <- geepack::geeglm(status ~ centre + treatment + sex + BL_status +
I(age-33) + I((age-33)^2),
data = df_resp_long,
family = binomial(link = "logit"),
id = subject,
waves = month,
corstr = "ar1",
scale.fix = TRUE)
summary(fit_geeglm_ar)
Call:
geepack::geeglm(formula = status ~ centre + treatment + sex +
BL_status + I(age - 33) + I((age - 33)^2), family = binomial(link = "logit"),
data = df_resp_long, id = subject, waves = month, corstr = "ar1",
scale.fix = TRUE)
Coefficients:
Estimate Std.err Wald Pr(>|W|)
(Intercept) -1.949751 0.451399 18.66 1.6e-05 ***
centre2 0.627506 0.375800 2.79 0.09496 .
treatmentTreatment 1.286413 0.377526 11.61 0.00066 ***
sexMale 0.388172 0.485677 0.64 0.42415
BL_statusGood 1.942647 0.374568 26.90 2.1e-07 ***
I(age - 33) -0.033646 0.014935 5.08 0.02427 *
I((age - 33)^2) 0.002306 0.000755 9.32 0.00227 **
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Correlation structure = ar1
Scale is fixed.
Link = identity
Estimated Correlation Parameters:
Estimate Std.err
alpha 0.428 0.0552
Number of clusters: 111 Maximum cluster size: 4 16.3.4 Paramgeter Estimates Table
apaSupp::tab_gees(list("Independent" = fit_geeglm_in,
"Exchangable" = fit_geeglm_ex,
"Autoregress" = fit_geeglm_ar),
narrow = TRUE)
| Independent | Exchangable | Autoregress | |||
|---|---|---|---|---|---|---|
OR | 95% CI | OR | 95% CI | OR | 95% CI | |
centre | ||||||
1 | — | — | — | — | — | — |
2 | 1.71 | [0.81, 3.59] | 1.71 | [0.81, 3.59] | 1.87 | [0.90, 3.91] |
treatment | ||||||
Placebo | — | — | — | — | — | — |
Treatment | 3.88 | [1.85, 8.14]*** | 3.88 | [1.85, 8.14]*** | 3.62 | [1.73, 7.59]*** |
sex | ||||||
Female | — | — | — | — | — | — |
Male | 1.53 | [0.59, 3.95] | 1.53 | [0.59, 3.95] | 1.47 | [0.57, 3.82] |
BL_status | ||||||
Poor | — | — | — | — | — | — |
Good | 6.82 | [3.25, 14.28]*** | 6.82 | [3.25, 14.28]*** | 6.98 | [3.35, 14.54]*** |
I(age - 33) | 0.96 | [0.94, 0.99]* | 0.96 | [0.94, 0.99]* | 0.97 | [0.94, 1.00]* |
I((age - 33)^2) | 1.00 | [1.00, 1.00]*** | 1.00 | [1.00, 1.00]*** | 1.00 | [1.00, 1.00]** |
Note. | ||||||
* p < .05. ** p < .01. *** p < .001. | ||||||
16.3.5 Compare Models
QIC
fit_geeglm_in 491
fit_geeglm_ex 491
fit_geeglm_ar 49216.3.6 Final Model
16.3.6.1 Estimates on both the logit and odds-ratio scales
Odds Ratio | Logit Scale | ||||||
|---|---|---|---|---|---|---|---|
OR | 95% CI | b | (SE) | p | |||
centre | |||||||
1 | — | — | — | — | |||
2 | 1.71 | [0.81, 3.59] | 0.53 | (0.38) | .159 | ||
treatment | |||||||
Placebo | — | — | — | — | |||
Treatment | 3.88 | [1.85, 8.14] | 1.36 | (0.38) | < .001*** | ||
sex | |||||||
Female | — | — | — | — | |||
Male | 1.53 | [0.59, 3.95] | 0.43 | (0.48) | .378 | ||
BL_status | |||||||
Poor | — | — | — | — | |||
Good | 6.82 | [3.25, 14.28] | 1.92 | (0.38) | < .001*** | ||
I(age - 33) | 0.96 | [0.94, 0.99] | -0.04 | (0.02) | .014* | ||
I((age - 33)^2) | 1.00 | [1.00, 1.00] | 0.00 | (0.00) | < .001*** | ||
(Intercept) | -1.97 | (0.45) | < .001*** | ||||
Note. N = 444 observations on 111 participants. Correlation structure: exchangeable | |||||||
* p < .05. ** p < .01. *** p < .001. | |||||||
16.3.6.2 Interpretation
centre: Controlling for baseline status, sex, age, and treatment, participants at the two centers did not significantly differ in respiratory status during the interventionsex: Controlling for baseline status, center, age, and treatment, a participant’s respiratory status did not differ between the two sexes.BL_status: Controlling for sex, center, age, and treatment, those with good baseline status had nearly 7 times higher odds of having a good respiratory status, compared to participants that starts out poor.age: Controlling for baseline status, sex, center, and treatment, the role of age was non-linear, such that the odds of a good respiratory status was lowest for patients age 40 and better for those that were either younger or older.
Most importantly:
treatment: Controlling for baseline status, sex, age, and center, those on the treatment had 3.88 time higher odds of having a good respiratory status, compared to similar participants that were randomized to the placebo.
16.3.7 Predicted Probabilities
16.3.7.1 Make predictions
What is the change a 40 year old man in poor condition at center 1 change of being rated as being in “Good” respiratory condition?
fit_geeglm_ex %>%
emmeans::emmeans(pairwise ~ treatment,
at = list(centre = "1",
sex = "Male",
age = 40,
BL_status = "Poor"),
type = "response")$emmeans
treatment prob SE df lower.CL upper.CL
Placebo 0.158 0.0548 437 0.0767 0.296
Treatment 0.421 0.1160 437 0.2211 0.650
Covariance estimate used: vbeta
Confidence level used: 0.95
Intervals are back-transformed from the logit scale
$contrasts
contrast odds.ratio SE df null t.ratio p.value
Placebo / Treatment 0.258 0.0973 437 1 -3.590 0.0004
Tests are performed on the log odds ratio scale A 40 year old man in poor condition at center 1 has a 15.8% change of being rated as being in “Good” respiratory condition if he was randomized to placebo.
A 40 year old man in poor condition at center 1 has a 42.1% change of being rated as being in “Good” respiratory condition if he was randomized to placebo.
The odds ratio for treatment is:
[1] 3.8716.3.8 Marginal Plot to Visualize the Model
16.3.8.1 Quickest Version
The interactions::interact_plot() function can only investigate 3 variables at once:
predthe x-axis (must be continuous)modxdifferent lines (may be categorical or continuous)mod2side-by-side panels (may be categorical or continuous)
All other variables must be held constant.
interactions::interact_plot(model = fit_geeglm_ex, # model name
pred = age, # x-axis
modx = treatment, # lines
mod2 = sex, # panels
data = df_resp_long,
at = list(centre = "1",
BL_status = "Good"), # hold constant
type = "mean_subject") +
theme_bw()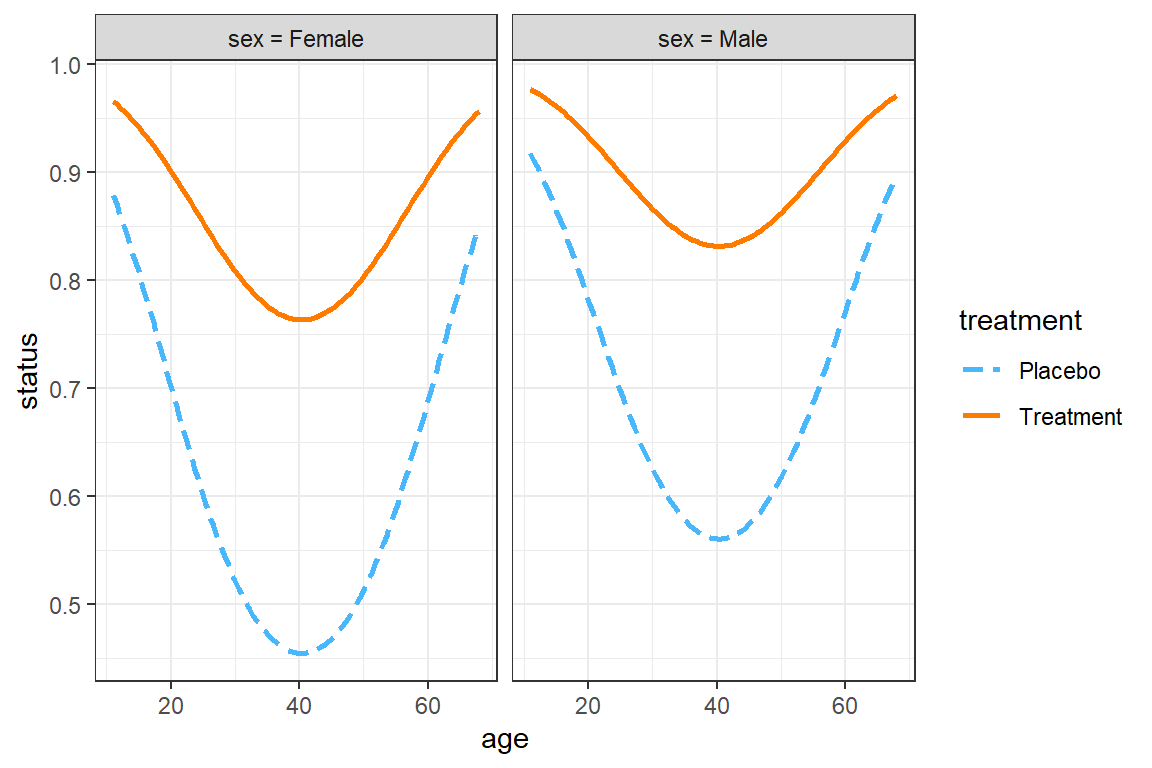
16.3.8.2 Full Version
Makes a dataset with all combinations
fit_geeglm_ex %>%
emmeans::emmeans(~ centre + treatment + sex + age + BL_status,
at = list(age = c(20, 35, 50)),
type = "response") %>%
data.frame() centre treatment sex age BL_status prob SE df lower.CL upper.CL
1 1 Placebo Female 20 Poor 0.257 0.0655 437 0.1494 0.404
2 2 Placebo Female 20 Poor 0.371 0.1211 437 0.1751 0.620
3 1 Treatment Female 20 Poor 0.572 0.0709 437 0.4310 0.703
4 2 Treatment Female 20 Poor 0.696 0.0901 437 0.4975 0.841
5 1 Placebo Male 20 Poor 0.346 0.1033 437 0.1773 0.565
6 2 Placebo Male 20 Poor 0.474 0.1190 437 0.2610 0.697
7 1 Treatment Male 20 Poor 0.672 0.1335 437 0.3840 0.871
8 2 Treatment Male 20 Poor 0.778 0.0996 437 0.5301 0.916
9 1 Placebo Female 35 Poor 0.116 0.0472 437 0.0504 0.245
10 2 Placebo Female 35 Poor 0.183 0.0916 437 0.0628 0.427
11 1 Treatment Female 35 Poor 0.337 0.0661 437 0.2214 0.476
12 2 Treatment Female 35 Poor 0.465 0.1104 437 0.2662 0.675
13 1 Placebo Male 35 Poor 0.167 0.0567 437 0.0827 0.309
14 2 Placebo Male 35 Poor 0.255 0.0845 437 0.1252 0.451
15 1 Treatment Male 35 Poor 0.438 0.1191 437 0.2313 0.669
16 2 Treatment Male 35 Poor 0.571 0.1126 437 0.3501 0.767
17 1 Placebo Female 50 Poor 0.134 0.0588 437 0.0539 0.295
18 2 Placebo Female 50 Poor 0.209 0.1013 437 0.0732 0.468
19 1 Treatment Female 50 Poor 0.375 0.0844 437 0.2281 0.549
20 2 Treatment Female 50 Poor 0.506 0.1102 437 0.3008 0.709
21 1 Placebo Male 50 Poor 0.191 0.0674 437 0.0913 0.358
22 2 Placebo Male 50 Poor 0.288 0.0862 437 0.1502 0.480
23 1 Treatment Male 50 Poor 0.479 0.1261 437 0.2539 0.713
24 2 Treatment Male 50 Poor 0.611 0.1030 437 0.4009 0.786
25 1 Placebo Female 20 Good 0.702 0.0736 437 0.5410 0.824
26 2 Placebo Female 20 Good 0.801 0.0626 437 0.6501 0.897
27 1 Treatment Female 20 Good 0.901 0.0408 437 0.7874 0.957
28 2 Treatment Female 20 Good 0.940 0.0245 437 0.8696 0.973
29 1 Placebo Male 20 Good 0.783 0.1022 437 0.5252 0.921
30 2 Placebo Male 20 Good 0.860 0.0612 437 0.6936 0.944
31 1 Treatment Male 20 Good 0.933 0.0499 437 0.7436 0.985
32 2 Treatment Male 20 Good 0.960 0.0268 437 0.8588 0.989
33 1 Placebo Female 35 Good 0.472 0.0943 437 0.2980 0.653
34 2 Placebo Female 35 Good 0.604 0.1031 437 0.3953 0.781
35 1 Treatment Female 35 Good 0.776 0.0650 437 0.6244 0.879
36 2 Treatment Female 35 Good 0.855 0.0442 437 0.7456 0.923
37 1 Placebo Male 35 Good 0.578 0.1205 437 0.3414 0.783
38 2 Placebo Male 35 Good 0.700 0.0824 437 0.5192 0.835
39 1 Treatment Male 35 Good 0.842 0.0878 437 0.5929 0.951
40 2 Treatment Male 35 Good 0.901 0.0481 437 0.7592 0.963
41 1 Placebo Female 50 Good 0.513 0.1108 437 0.3057 0.716
42 2 Placebo Female 50 Good 0.643 0.1013 437 0.4303 0.811
43 1 Treatment Female 50 Good 0.803 0.0689 437 0.6343 0.906
44 2 Treatment Female 50 Good 0.875 0.0401 437 0.7728 0.935
45 1 Placebo Male 50 Good 0.617 0.1242 437 0.3646 0.819
46 2 Placebo Male 50 Good 0.734 0.0736 437 0.5678 0.852
47 1 Treatment Male 50 Good 0.862 0.0808 437 0.6219 0.960
48 2 Treatment Male 50 Good 0.914 0.0409 437 0.7927 0.968In order to include all FIVE variables, we must do it the LONG way…
fit_geeglm_ex %>%
emmeans::emmeans(~ centre + treatment + sex + age + BL_status,
at = list(age = seq(from = 11, to = 68, by = 1)),
type = "response",
level = .68) %>%
data.frame() %>%
ggplot(aes(x = age,
y = prob,
group = interaction(sex, treatment))) +
geom_ribbon(aes(ymin = lower.CL,
ymax = upper.CL,
fill = fct_rev(sex)),
alpha = .3) +
geom_line(aes(color = fct_rev(sex),
linetype = fct_rev(treatment))) +
theme_bw() +
facet_grid(centre ~ BL_status, labeller = label_both) +
labs(x = "Age, years",
y = "Predicted Probability of GOOD Respiratory Status",
color = "Sex:",
fill = "Sex:",
linetype = "Assignment:") +
theme(legend.position = "bottom")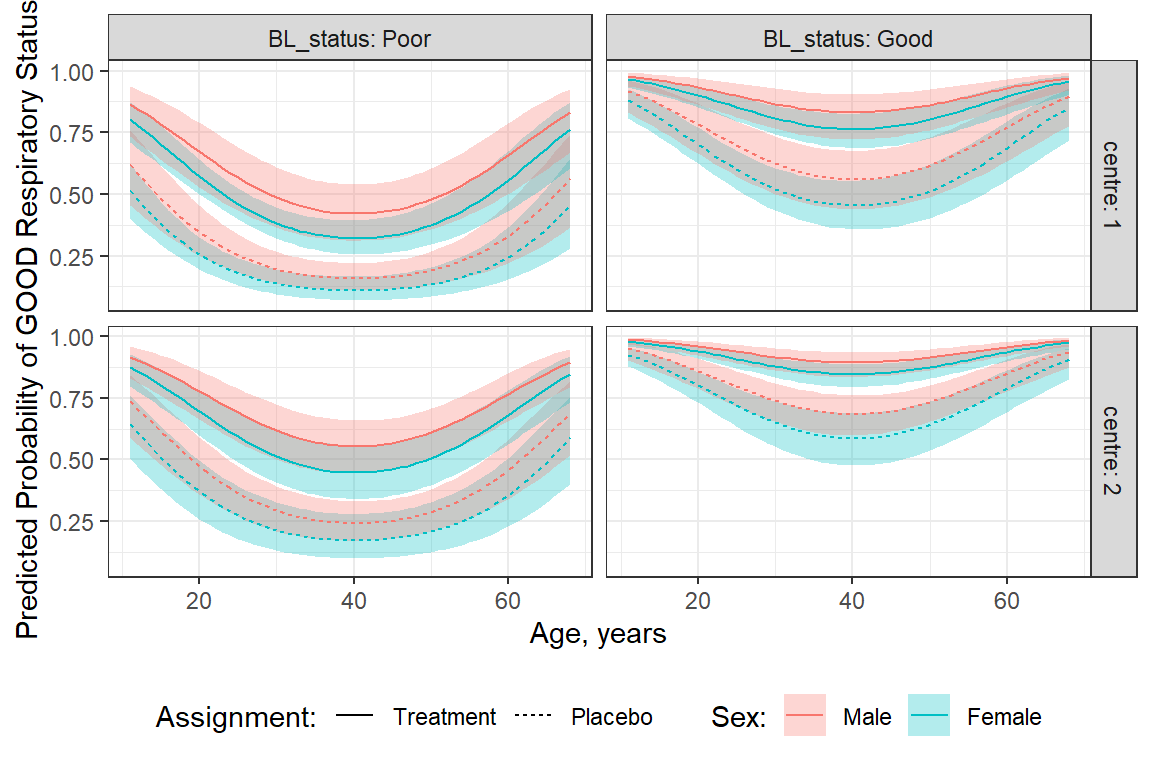
16.3.8.3 Females in Center 1
This example uses default settings.
interactions::interact_plot(model = fit_geeglm_ex,
pred = age,
modx = treatment,
mod2 = BL_status) +
theme_bw()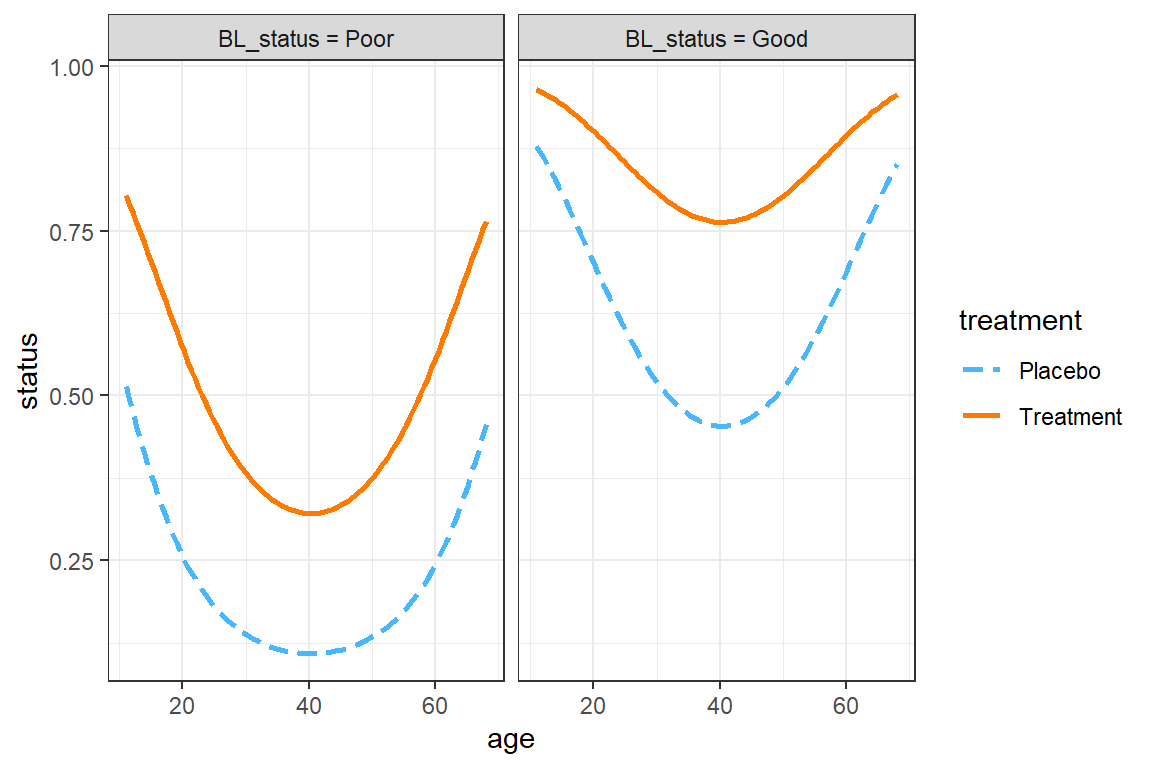
interactions::interact_plot(model = fit_geeglm_ex,
pred = age,
modx = treatment,
mod2 = BL_status,
at = list(sex = "Female",
centre = "1")) +
theme_bw()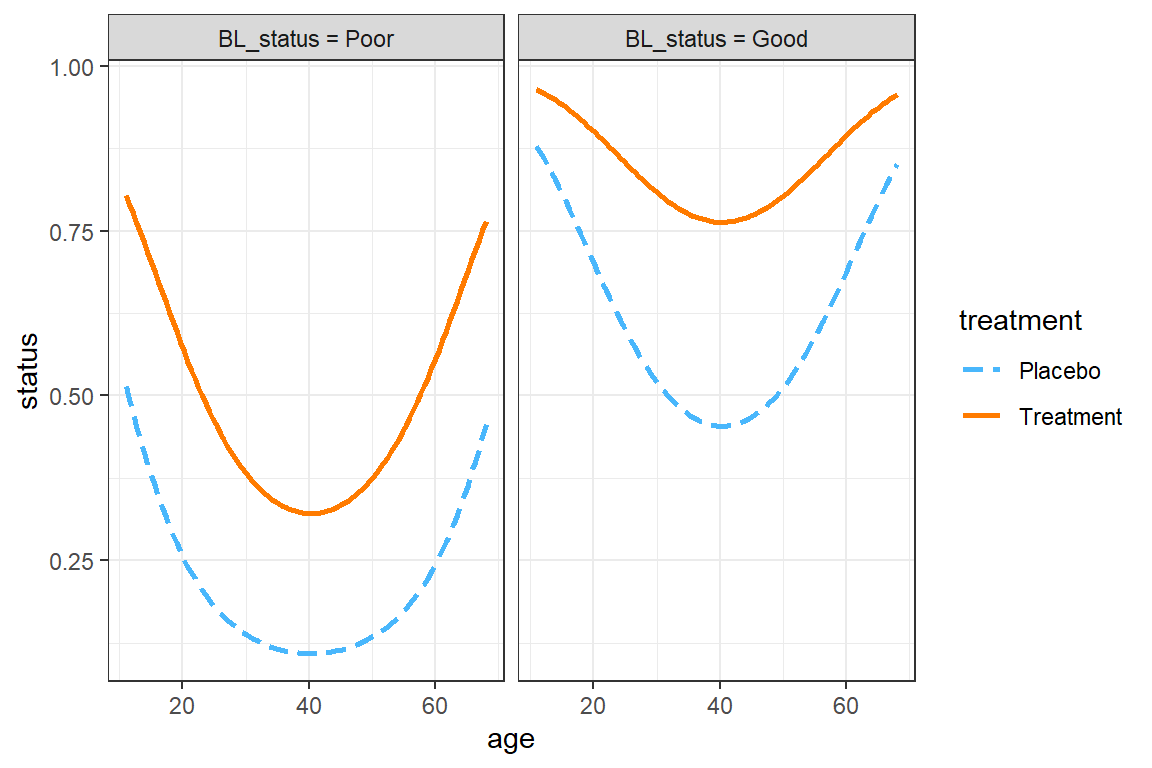
16.3.8.4 Males in Center 2
This example is more preped for publication.
interactions::interact_plot(model = fit_geeglm_ex,
pred = age,
modx = treatment,
mod2 = BL_status,
at = list(sex = "Male",
centre = "2"),
x.label = "Age in Years",
y.label = "Predicted Probability of 'Good' Respiratory Status",
legend.main = "Intervention: ",
mod2.labels = c("Poor at Baseline",
"Good at Baseline"),
colors = rep("black", times = 2)) +
theme_bw() +
theme(legend.position = c(1, 0),
legend.justification = c(1.1, -0.1),
legend.background = element_rect(color = "black"),
legend.key.width = unit(1.5, "cm")) +
labs(caption = "Note: Probibilities shown are specific to males at center 2")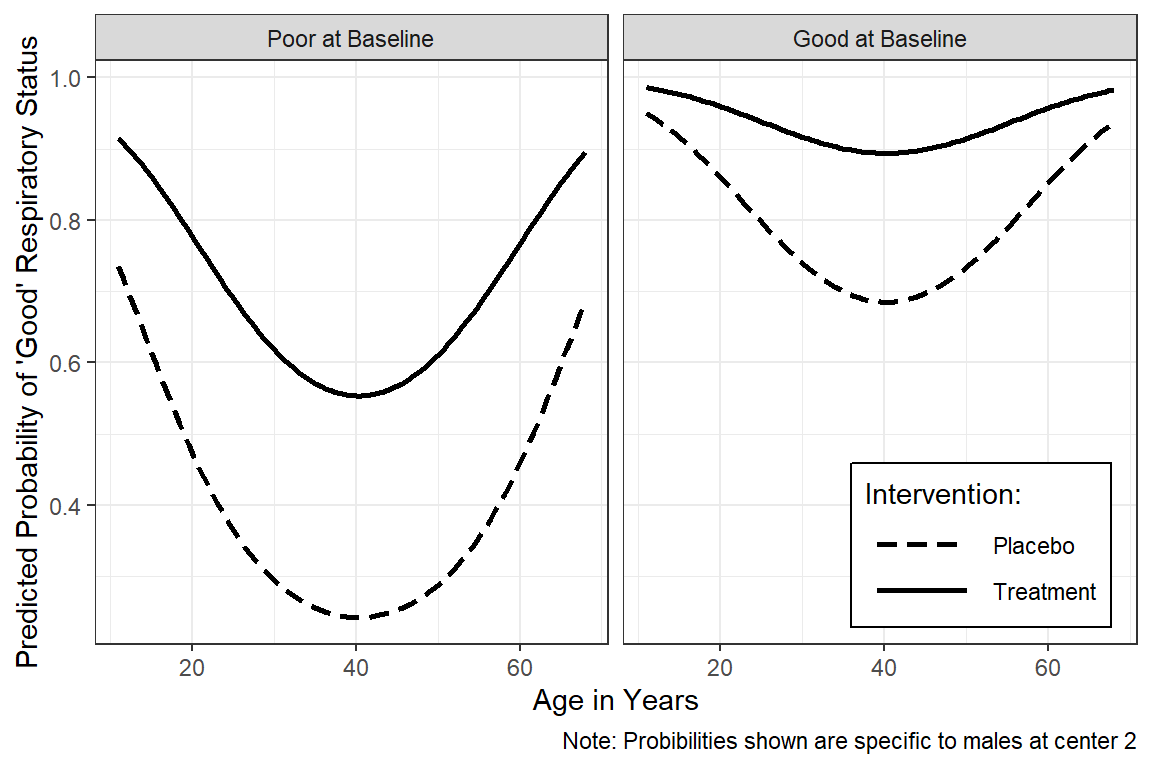
fit_geeglm_ex %>%
emmeans::emmeans(~ centre + treatment + sex + age + BL_status,
at = list(age = seq(from = 11, to = 68, by = 1),
sex = "Male",
centre = "2"),
type = "response",
level = .68) %>%
data.frame() %>%
ggplot(aes(x = age,
y = prob,
group = interaction(sex, treatment))) +
geom_ribbon(aes(ymin = lower.CL,
ymax = upper.CL),
alpha = .2) +
geom_line(aes(linetype = fct_rev(treatment))) +
theme_bw() +
facet_grid(~ BL_status) +
labs(x = "Age, years",
y = "Predicted Probability of\nGOOD Respiratory Status",
color = "Sex:",
fill = "Sex:",
linetype = "Assignment:") +
theme(legend.position = "bottom")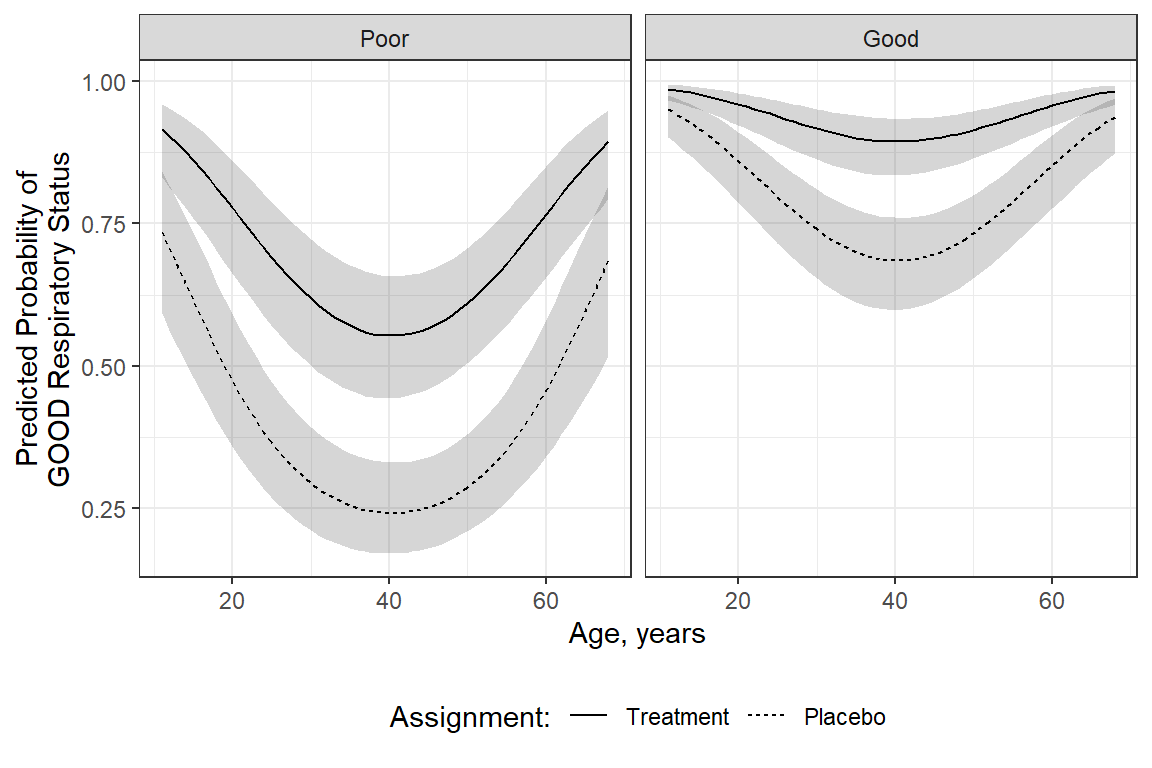
16.4 Conclusion
The Research Question
The question of interest is to assess whether the treatment is effective and to estimate its effect.
The Conclusion
After accounting for baseline status, age, sex and center, participants in the active treatment group had nearly four times higher odds of having ‘good’ respiratory status, when compared to the placebo, exp(b) = 3.881, p<.001, 95% CI [1.85, 8.14].