23 GLMM, Count Outcome: Bolous
GLMM FAQ by: Ben Bolker and others
23.1 Packages
23.1.1 CRAN
library(tidyverse) # all things tidy
library(pander) # nice looking genderal tabulations
library(furniture) # nice table1() descriptives
library(texreg) # Convert Regression Output to LaTeX or HTML Tables
library(psych) # contains some useful functions, like headTail
library(lme4) # Linear, generalized linear, & nonlinear mixed models
library(gee) # Generalized Estimating Equations
library(optimx) # Optimizers for use in lme4::glmer
library(MuMIn) # Multi-Model Inference (caluclate QIC)
library(effects) # Plotting estimated marginal means
library(interactions)
library(performance)
library(patchwork) # combining graphics\
library(cold) # this package holds the dataset we need23.2 Data Prep
Bolus data from Weiss 2005
Patient controlled analgesia (PCA) comparing 2 dosing regimes to self-control pain
The dataset has the number of requests per interval in 12 successive four-hourly intervals following abdominal surgery for 65 patients in a clinical trial to compare two groups (bolus/lock-out combinations).
Bolus= Large dose of medication given (usually intravenously by direct infusion injection or gravity drip) to raise blood-level concentrations to a therapeutic level
A ‘lock-out’ period followed each dose, where subject may not administer more medication.
- Lock-out time is twice as long in 2mg/dose group
- Allows for max of 30 dosages in 2mg/dose and 60 in 1mg/dose group in any 4-hour period
- No responses neared these upper limits
Variable List:
Indicators
idunique subject indicatortime11 consecutive 4-hour periods: 0, 1, 2, …, 10Outcome or dependent variable
count# doses recorded for in that 4-hour periodMain predictor or independent variable of interest
group1mg/dose group, 2mg/dose group
23.2.1 Import
'data.frame': 780 obs. of 4 variables:
$ id : int 1 1 1 1 1 1 1 1 1 1 ...
$ group: Factor w/ 2 levels "1mg","2mg": 2 2 2 2 2 2 2 2 2 2 ...
$ time : int 1 2 3 4 5 6 7 8 9 10 ...
$ y : int 5 2 2 5 2 4 0 2 4 4 ... id group time y
1 1 2mg 1 5
2 1 2mg 2 2
3 1 2mg 3 2
4 1 2mg 4 5
... ... <NA> ... ...
777 65 1mg 9 6
778 65 1mg 10 1
779 65 1mg 11 2
780 65 1mg 12 023.2.2 Long Format
df_long <- df_raw %>%
dplyr::rename(count = y) %>%
dplyr::mutate(id = factor(id)) %>%
dplyr::mutate(time = time - 1) %>%
dplyr::mutate(timeF = factor(time)) %>%
dplyr::mutate(group = factor(group))'data.frame': 780 obs. of 5 variables:
$ id : Factor w/ 65 levels "1","2","3","4",..: 1 1 1 1 1 1 1 1 1 1 ...
$ group: Factor w/ 2 levels "1mg","2mg": 2 2 2 2 2 2 2 2 2 2 ...
$ time : num 0 1 2 3 4 5 6 7 8 9 ...
$ count: int 5 2 2 5 2 4 0 2 4 4 ...
$ timeF: Factor w/ 12 levels "0","1","2","3",..: 1 2 3 4 5 6 7 8 9 10 ... id group time count timeF
1 1 2mg 0 5 0
2 1 2mg 1 2 1
3 1 2mg 2 2 2
4 1 2mg 3 5 3
... <NA> <NA> ... ... <NA>
777 65 1mg 8 6 8
778 65 1mg 9 1 9
779 65 1mg 10 2 10
780 65 1mg 11 0 1123.2.3 Wide Format
df_wide <- df_long %>%
dplyr::select(-timeF) %>%
tidyr::pivot_wider(names_from = time,
names_prefix = "time_",
values_from = count)tibble [65 × 14] (S3: tbl_df/tbl/data.frame)
$ id : Factor w/ 65 levels "1","2","3","4",..: 1 2 3 4 5 6 7 8 9 10 ...
$ group : Factor w/ 2 levels "1mg","2mg": 2 2 2 2 2 2 2 2 2 2 ...
$ time_0 : int [1:65] 5 5 6 4 8 10 2 14 7 3 ...
$ time_1 : int [1:65] 2 3 4 1 7 6 4 2 10 0 ...
$ time_2 : int [1:65] 2 5 8 4 7 5 7 10 9 7 ...
$ time_3 : int [1:65] 5 4 4 3 6 4 4 8 8 1 ...
$ time_4 : int [1:65] 2 4 3 3 8 8 8 17 9 9 ...
$ time_5 : int [1:65] 4 5 12 3 6 7 4 4 4 8 ...
$ time_6 : int [1:65] 0 0 1 1 5 4 4 2 2 0 ...
$ time_7 : int [1:65] 2 6 0 7 4 6 3 6 6 1 ...
$ time_8 : int [1:65] 4 3 6 5 7 5 1 8 1 5 ...
$ time_9 : int [1:65] 4 2 5 0 2 2 3 9 8 4 ...
$ time_10: int [1:65] 4 7 3 1 5 4 4 1 5 1 ...
$ time_11: int [1:65] 2 4 5 1 0 4 6 4 5 0 ... id group time_0 time_1 time_2 time_3 time_4 time_5 time_6 time_7 time_8
1 1 2mg 5 2 2 5 2 4 0 2 4
2 2 2mg 5 3 5 4 4 5 0 6 3
3 3 2mg 6 4 8 4 3 12 1 0 6
4 4 2mg 4 1 4 3 3 3 1 7 5
5 <NA> <NA> ... ... ... ... ... ... ... ... ...
6 62 1mg 25 0 13 7 9 19 4 5 11
7 63 1mg 12 11 7 6 9 3 4 6 6
8 64 1mg 3 8 4 22 11 16 4 5 9
9 65 1mg 14 2 2 2 4 4 3 7 6
time_9 time_10 time_11
1 4 4 2
2 2 7 4
3 5 3 5
4 0 1 1
5 ... ... ...
6 10 6 9
7 6 6 0
8 12 12 7
9 1 2 023.3 Exploratory Data Analysis
23.3.1 Summary Statistics
23.3.1.1 Demographics and Baseline
| Total | 1mg | 2mg | p-value |
|---|---|---|---|---|
Count at Baseline | 9.77 (6.84) | 10.17 (7.69) | 9.30 (5.78) | .605 |
Note. Continuous variables are summarized with means (SD) and significant group differences assessed via independent t-tests. Categorical variables are summarized with counts (%) and significant group differences assessed via Chi-squared tests for independence. | ||||
* p < .05. ** p < .01. *** p < .001. | ||||
23.3.1.2 Status over Time
df_wide %>%
dplyr::select(group,
time_0,
time_1, time_2, time_3, time_4, time_5,
time_6, time_7, time_8, time_9, time_10,
time_11) %>%
apaSupp::tab1(split = "group")
| Total | 1mg | 2mg | p-value |
|---|---|---|---|---|
time_0 | 9.77 (6.84) | 10.17 (7.69) | 9.30 (5.78) | .605 |
time_1 | 6.03 (5.17) | 6.49 (5.58) | 5.50 (4.70) | .442 |
time_2 | 6.74 (6.37) | 7.86 (7.75) | 5.43 (4.00) | .112 |
time_3 | 7.37 (6.25) | 9.29 (7.12) | 5.13 (4.13) | .005** |
time_4 | 8.68 (5.94) | 9.63 (6.56) | 7.57 (5.01) | .156 |
time_5 | 6.43 (4.74) | 7.37 (5.54) | 5.33 (3.36) | .074 |
time_6 | 5.37 (4.80) | 6.60 (5.00) | 3.93 (4.19) | .022* |
time_7 | 4.82 (4.16) | 5.74 (4.78) | 3.73 (3.04) | .045* |
time_8 | 4.77 (3.62) | 4.91 (4.01) | 4.60 (3.16) | .725 |
time_9 | 5.69 (4.75) | 6.34 (5.51) | 4.93 (3.63) | .222 |
time_10 | 4.98 (4.43) | 6.26 (5.45) | 3.50 (2.06) | .008** |
time_11 | 4.77 (4.80) | 5.89 (5.81) | 3.47 (2.83) | .034* |
Note. Continuous variables are summarized with means (SD) and significant group differences assessed via independent t-tests. Categorical variables are summarized with counts (%) and significant group differences assessed via Chi-squared tests for independence. | ||||
* p < .05. ** p < .01. *** p < .001. | ||||
23.3.2 Visualize
23.3.2.1 Person Profile Plots, by Group
df_long %>%
ggplot(aes(x = time,
y = count)) +
geom_line(aes(group = id),
color = "black",
alpha = .4) +
facet_grid(. ~ group) +
theme_bw()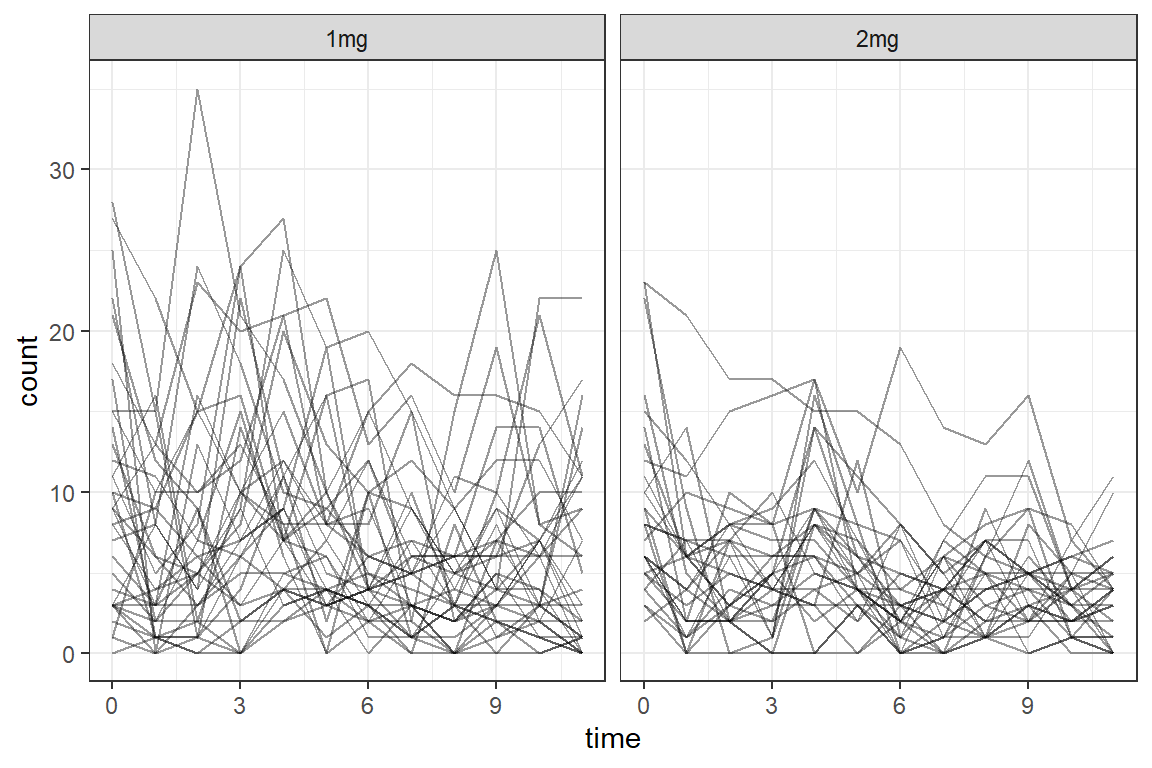
df_long %>%
ggplot(aes(x = time,
y = count)) +
geom_smooth(aes(group = id),
method = "lm",
color = "gray",
se = FALSE) +
facet_grid(. ~ group) +
theme_bw()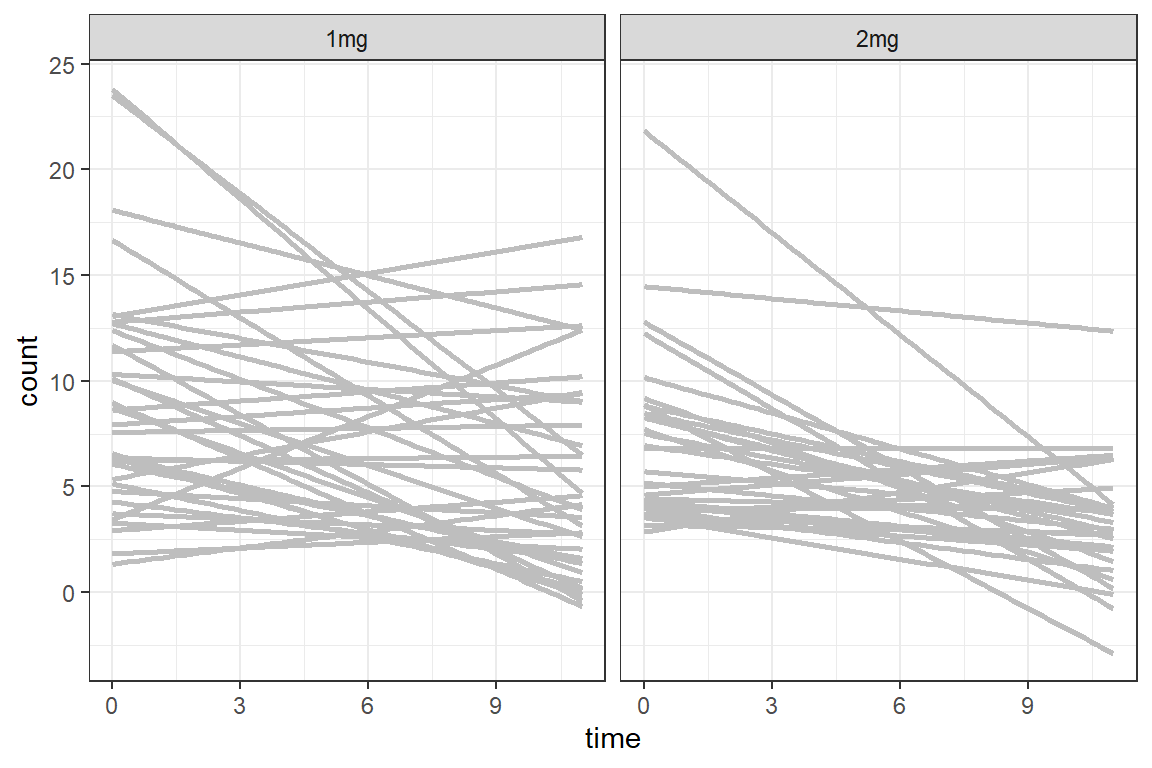
df_long %>%
ggplot(aes(x = time,
y = count)) +
geom_smooth(aes(group = id),
method = "lm",
formula = y ~ poly(x, 2),
color = "gray",
se = FALSE) +
facet_grid(. ~ group) +
theme_bw()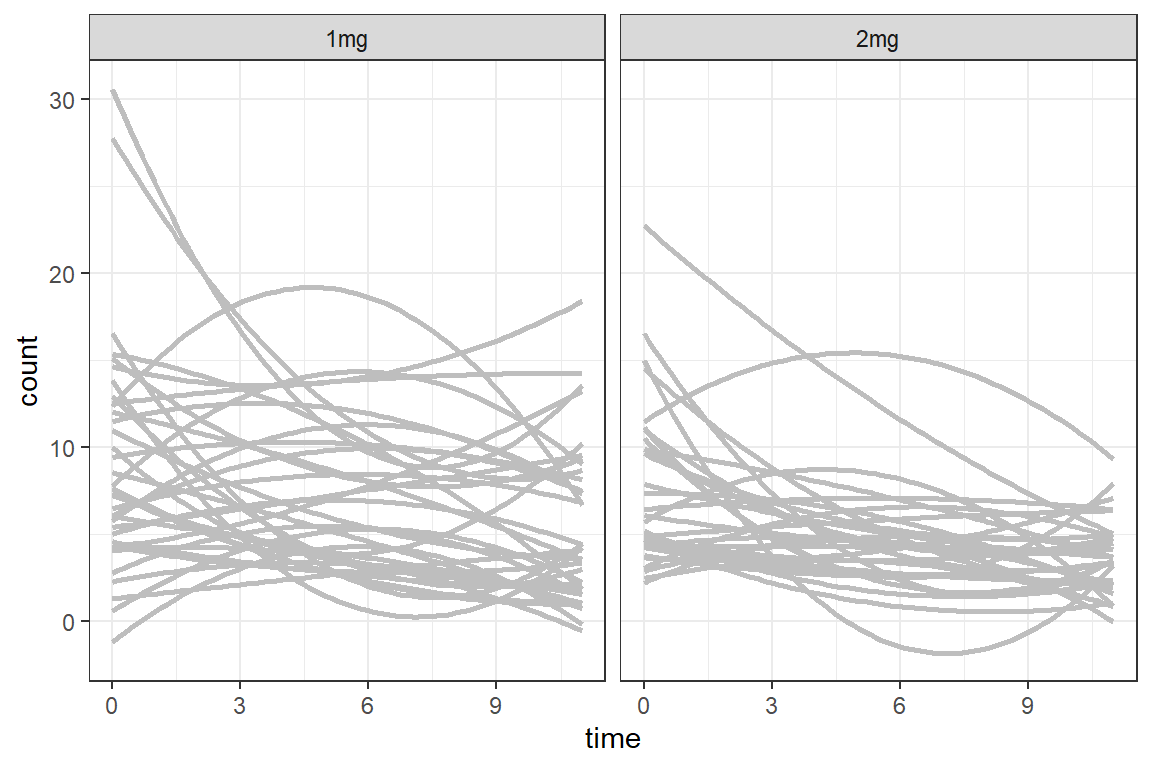
df_long %>%
ggplot(aes(x = time,
y = count)) +
geom_line(aes(group = id)) +
geom_smooth(method = "loess",
color = "blue",
se = FALSE) +
geom_smooth(method = "lm",
color = "red",
se = FALSE) +
facet_grid(. ~ group) +
theme_bw()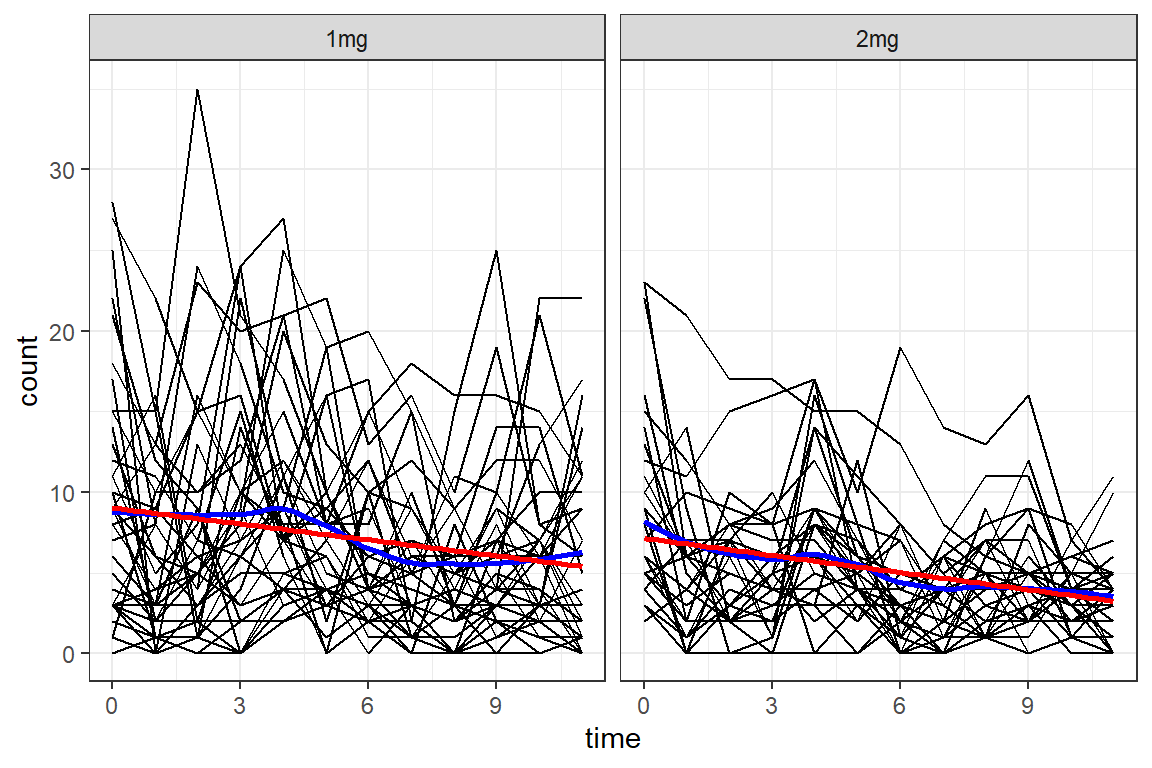
df_long %>%
ggplot(aes(x = time,
y = log(count + .1))) +
geom_line(aes(group = id)) +
geom_smooth(method = "loess",
color = "blue",
se = FALSE) +
geom_smooth(method = "lm",
color = "red",
se = FALSE) +
facet_grid(. ~ group) +
theme_bw()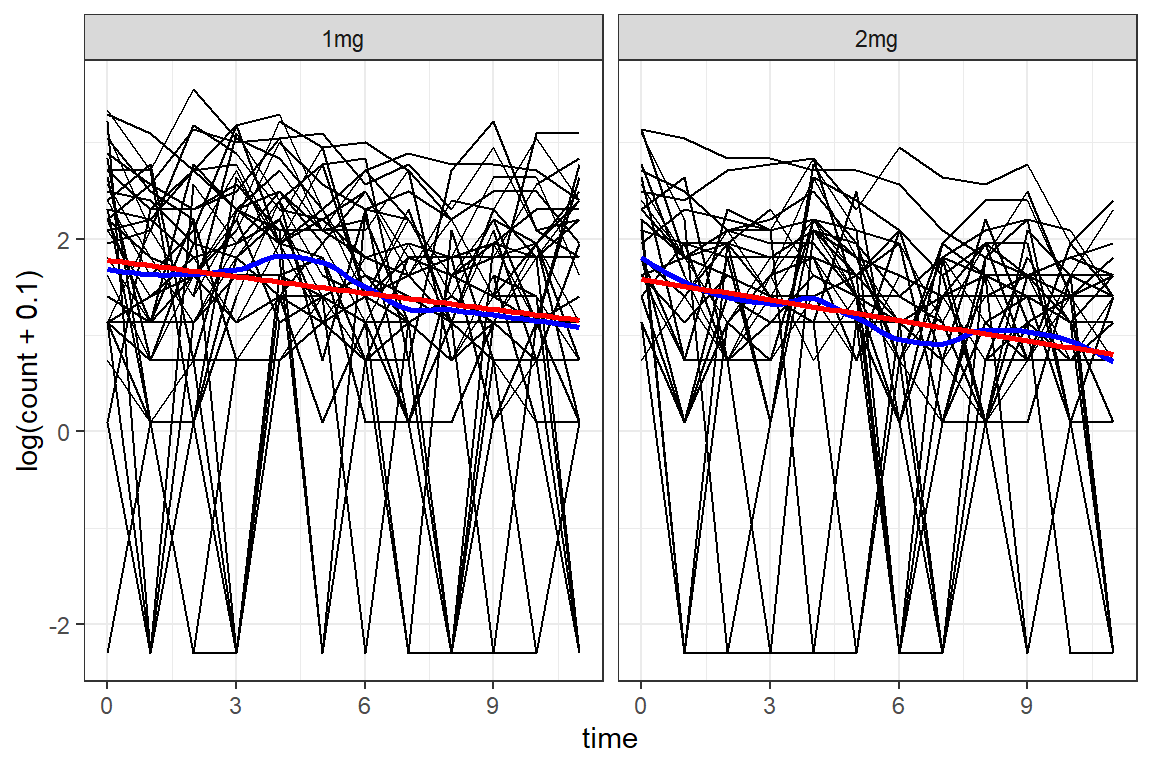
df_long %>%
ggplot(aes(x = time,
y = count)) +
geom_smooth(aes(color = "1. Loess"),
method = "loess",
se = FALSE) +
geom_smooth(aes(color = "2. Linear"),
method = "lm",
se = FALSE) +
geom_smooth(aes(color = "3. Quadratic"),
method = "lm",
formula = y ~ poly(x, 2),
se = FALSE) +
geom_smooth(aes(color = "4. Cubic"),
method = "lm",
formula = y ~ poly(x, 3),
se = FALSE) +
facet_grid(. ~ group) +
theme_bw() +
labs(color = "Type of Smoother") +
theme(legend.position = "bottom",
legend.key.width = unit(1.5, "cm"))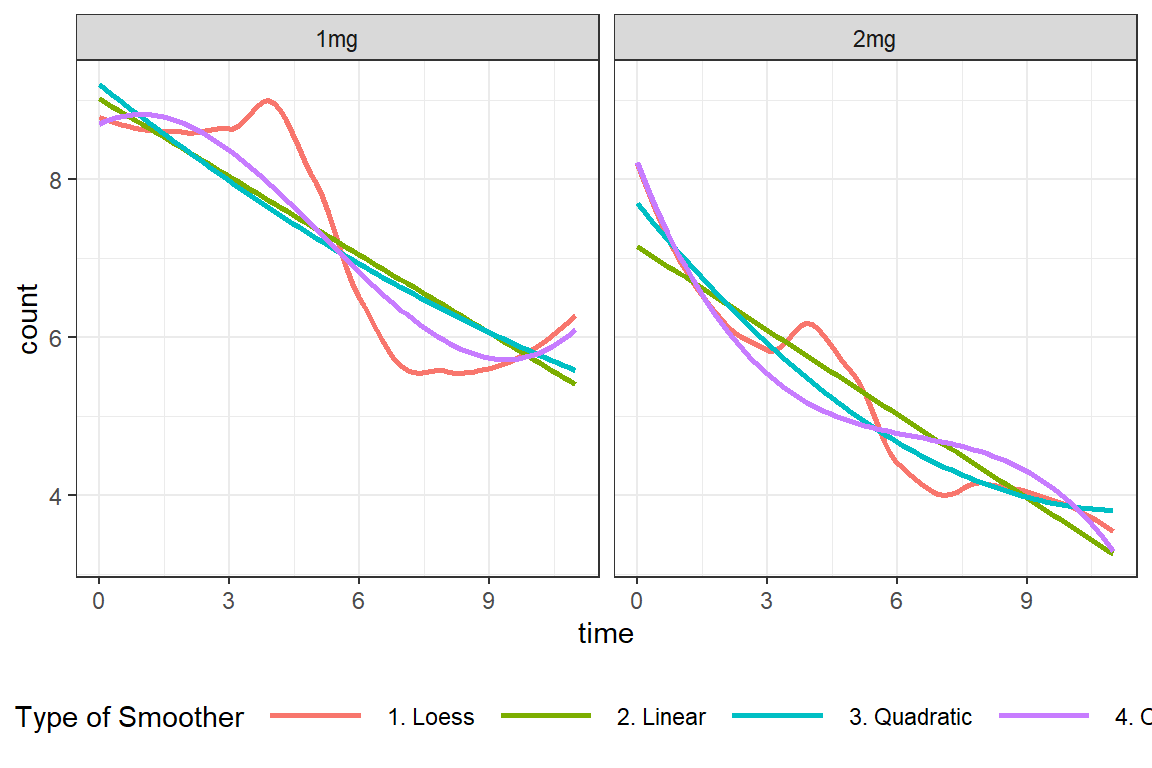
23.3.2.2 Means over Time, BY Group
df_long %>%
dplyr::group_by(group, timeF) %>%
dplyr::summarise(M = mean(count)) %>%
ggplot(aes(x = timeF,
y = M,
group = group,
color = group)) +
geom_point() +
geom_line() +
theme_bw() +
labs(x = "Four-week Intervals",
y = "Sample Mean Count",
color = "Dose") 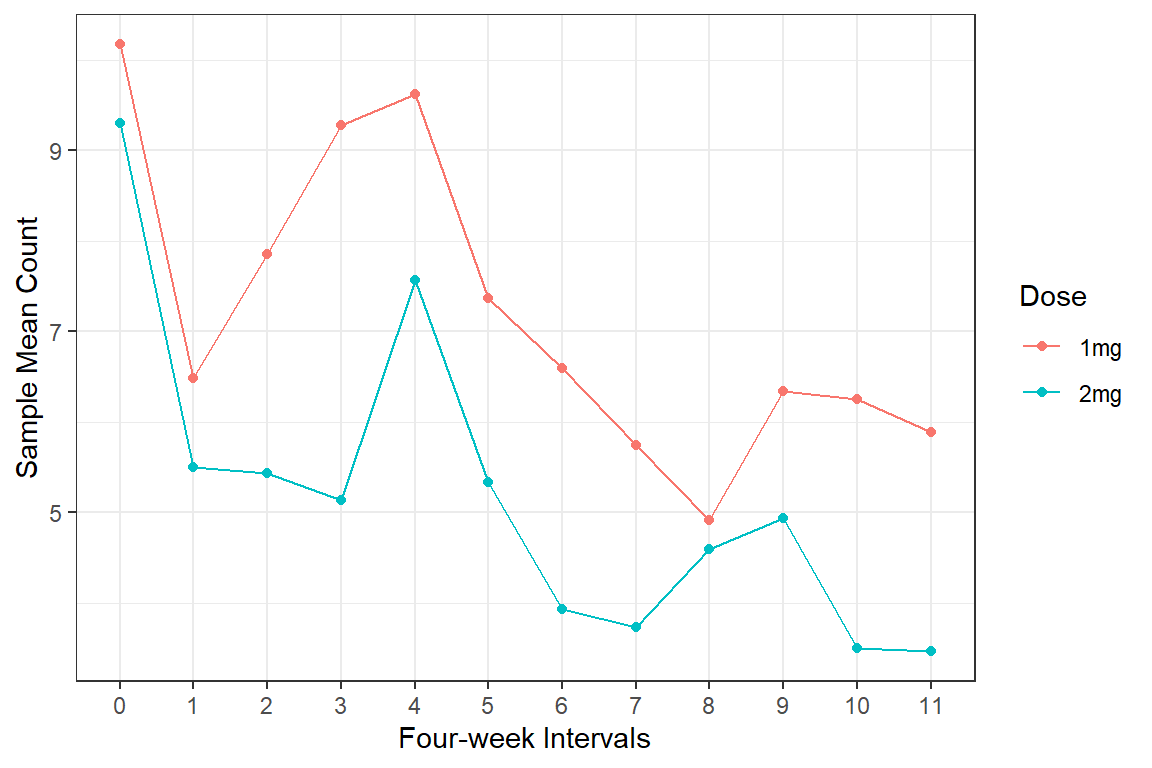
df_long %>%
dplyr::group_by(group, timeF) %>%
dplyr::summarise(M = mean(log(count + .1))) %>%
ggplot(aes(x = timeF,
y = M,
group = group,
color = group)) +
geom_point() +
geom_line() +
theme_bw() +
labs(x = "Four-week Intervals",
y = "Sample Mean Count",
color = "Dose") 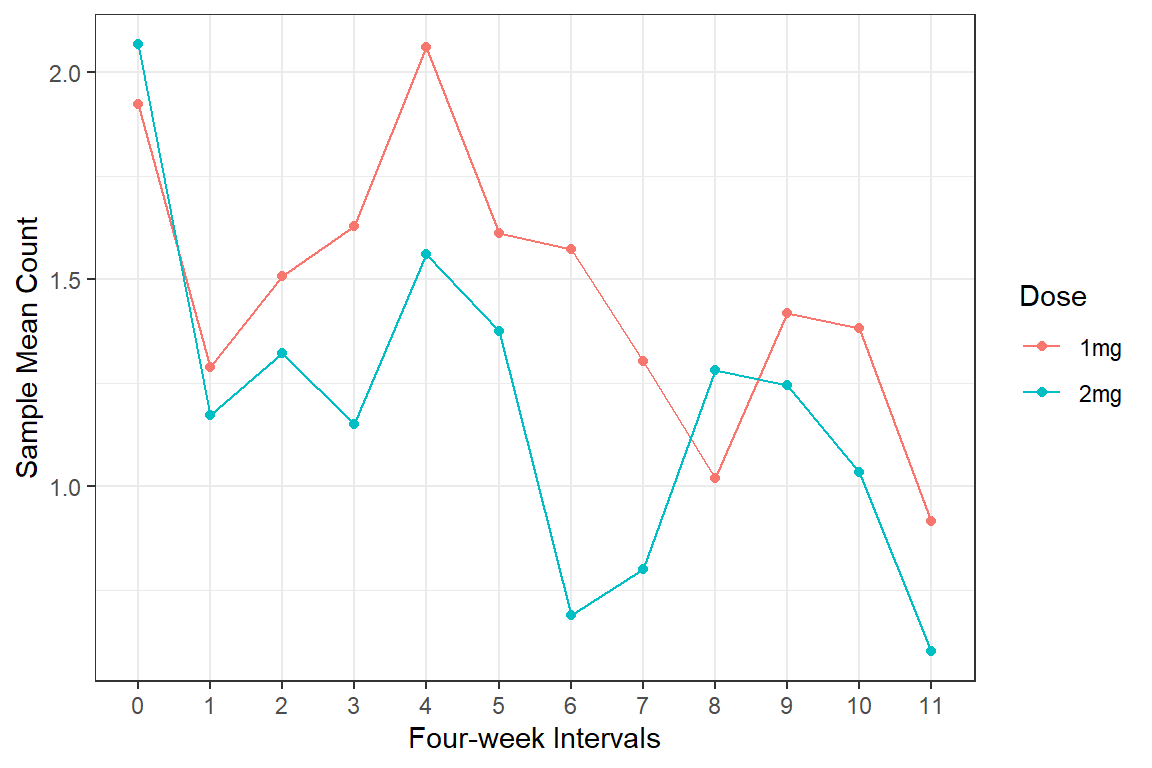
23.4 GEE Analysis
23.4.1 Fit Various Correlation Structures
mod_gee_ex <- geepack::geeglm(count ~ group + time + I(time^2),
id = id,
data = df_long,
family = poisson(link = "log"),
corstr = 'exchangeable')
mod_gee_ar <- geepack::geeglm(count ~ group + time + I(time^2),
id = id,
data = df_long,
family = poisson(link = "log"),
corstr = "ar1")
mod_gee_un <- geepack::geeglm(count ~ group + time + I(time^2),
id = id,
data = df_long,
family = poisson(link = "log"),
corstr = "unstructured") 23.4.2 Compare Corelation Structures
23.4.2.1 QIC: Model Fit
MuMIn::model.sel(mod_gee_ex,
mod_gee_ar,
mod_gee_un,
rank = "QIC") #sorts the best to the TOP, uses QIC(I) to choose corelation structureModel selection table
(Intrc) group time time^2 corstr qLik QIC delta weight
mod_gee_ex 2.26 + -0.0714 1.60e-03 exchng 4257 -8491 0.00 0.583
mod_gee_ar 2.39 + -0.1307 6.20e-03 ar1 4247 -8490 1.05 0.346
mod_gee_un 2.14 + -0.0646 -8.25e-06 unstrc 4202 -8487 4.22 0.071
Abbreviations:
corstr: exchng = 'exchangeable', unstrc = 'unstructured'
Models ranked by QIC(x) # Comparison of Model Performance Indices
Name | Model | RMSE | Sigma | Score_log | Score_spherical | AIC weights
-------------------------------------------------------------------------------
mod_gee_ex | geeglm | 5.207 | 2.017 | -3.617 | 0.028 | 1.000
mod_gee_ar | geeglm | 5.227 | 2.025 | -3.630 | 0.028 | 6.36e-05
mod_gee_un | geeglm | 5.283 | 2.210 | -3.687 | 0.028 | 2.16e-24
Name | AICc weights | BIC weights | Performance-Score
-----------------------------------------------------------
mod_gee_ex | 1.000 | 1.000 | 88.02%
mod_gee_ar | 6.36e-05 | 6.36e-05 | 35.84%
mod_gee_un | 2.16e-24 | 2.16e-24 | 14.29%23.4.2.2 Model Parameter Table
apaSupp::tab_gees(list("Exchangable" = mod_gee_ex,
"AutoRegressive" = mod_gee_ar,
"Unstructured" = mod_gee_un),
narrow = TRUE,
caption = "GEE: Main Effects Only, with Quadratic Time")
| Exchangable | AutoRegressive | Unstructured | |||
|---|---|---|---|---|---|---|
IRR | 95% CI | IRR | 95% CI | IRR | 95% CI | |
(Intercept) | 9.61 | [7.74, 11.94]*** | 10.88 | [8.81, 13.43]*** | 8.47 | [6.67, 10.74]*** |
group | ||||||
1mg | — | — | — | — | — | — |
2mg | 0.76 | [0.59, 0.98]* | 0.76 | [0.60, 0.98]* | 0.77 | [0.58, 1.02] |
time | 0.93 | [0.88, 0.99]* | 0.88 | [0.82, 0.94]*** | 0.94 | [0.87, 1.01] |
I(time^2) | 1.00 | [1.00, 1.01] | 1.01 | [1.00, 1.01]* | 1.00 | [0.99, 1.01] |
Note. | ||||||
* p < .05. ** p < .01. *** p < .001. | ||||||
23.4.3 Add Interaction Terms
mod_gee_ex2 <- geepack::geeglm(count ~ group + time,
id = id,
data = df_long,
family = poisson(link = "log"),
corstr = "exchangeable")
mod_gee_ex3 <- geepack::geeglm(count ~ group*time + group*I(time^2),
id = id,
data = df_long,
family = poisson(link = "log"),
corstr = "exchangeable") QIC
mod_gee_ex -8488
mod_gee_ex2 -8488
mod_gee_ex3 -8496apaSupp::tab_gees(list("Quadratic" = mod_gee_ex,
"Linear" = mod_gee_ex2,
"Interaction" = mod_gee_ex3),
narrow = TRUE,
caption = "GEE: Exchangeable")
| Quadratic | Linear | Interaction | |||
|---|---|---|---|---|---|---|
IRR | 95% CI | IRR | 95% CI | IRR | 95% CI | |
(Intercept) | 9.61 | [7.74, 11.94]*** | 9.38 | [7.73, 11.37]*** | 9.22 | [7.20, 11.79]*** |
group | ||||||
1mg | — | — | — | — | — | — |
2mg | 0.76 | [0.59, 0.98]* | 0.76 | [0.59, 0.98]* | 0.84 | [0.60, 1.18] |
time | 0.93 | [0.88, 0.99]* | 0.95 | [0.93, 0.97]*** | 0.95 | [0.88, 1.03] |
I(time^2) | 1.00 | [1.00, 1.01] | 1.00 | [0.99, 1.01] | ||
group * time | ||||||
2mg * time | 0.95 | [0.84, 1.06] | ||||
group * I(time^2) | ||||||
2mg * I(time^2) | 1.00 | [0.99, 1.01] | ||||
Note. | ||||||
* p < .05. ** p < .01. *** p < .001. | ||||||
23.4.4 Visualize Best Model
23.4.4.1 Model Parameter Table
Incident Rate Ratio | Log Scale | ||||||
|---|---|---|---|---|---|---|---|
IRR | 95% CI | b | (SE) | p | |||
(Intercept) | 9.38 | [7.73, 11.37] | 2.24 | (0.10) | < .001*** | ||
group | |||||||
1mg | — | — | — | — | |||
2mg | 0.76 | [0.59, 0.98] | -0.28 | (0.13) | .037* | ||
time | 0.95 | [0.93, 0.97] | -0.05 | (0.01) | < .001*** | ||
Note. N = 780 observations on 65 participants. Correlation structure: exchangeable | |||||||
* p < .05. ** p < .01. *** p < .001. | |||||||
23.4.4.2 Predict over a Grid
Estimated Marginal Mean Counts
mod_gee_ex2 %>%
emmeans::emmeans(pairwise ~group|time,
at = list(time = c(0, 5, 10)),
type = "response")$emmeans
time = 0:
group rate SE df lower.CL upper.CL
1mg 9.38 0.925 777 7.72 11.38
2mg 7.12 0.729 777 5.82 8.70
time = 5:
group rate SE df lower.CL upper.CL
1mg 7.14 0.675 777 5.93 8.59
2mg 5.42 0.497 777 4.52 6.49
time = 10:
group rate SE df lower.CL upper.CL
1mg 5.43 0.624 777 4.34 6.80
2mg 4.12 0.440 777 3.34 5.08
Covariance estimate used: vbeta
Confidence level used: 0.95
Intervals are back-transformed from the log scale
$contrasts
time = 0:
contrast ratio SE df null t.ratio p.value
1mg / 2mg 1.32 0.174 777 1 2.085 0.0374
time = 5:
contrast ratio SE df null t.ratio p.value
1mg / 2mg 1.32 0.174 777 1 2.085 0.0374
time = 10:
contrast ratio SE df null t.ratio p.value
1mg / 2mg 1.32 0.174 777 1 2.085 0.0374
Tests are performed on the log scale 23.4.4.3 Plot Estimated Marginal Means
interactions::interact_plot(model = mod_gee_ex2,
pred = time,
modx = group) # Can NOT do interval ribbons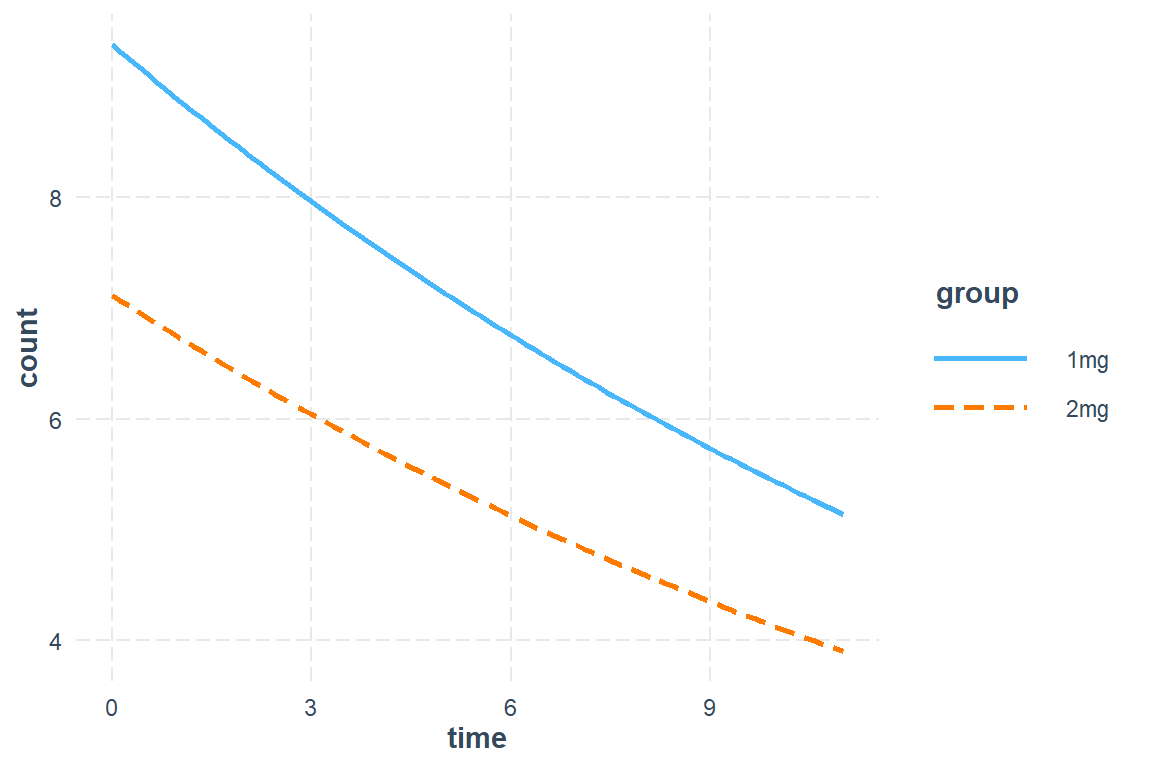
mod_gee_ex2 %>%
emmeans::emmeans(~group|time,
at = list(time = seq(from = 0, to = 11, by = .1)),
type = "response",
level = .685) %>%
data.frame() %>%
ggplot(aes(x = time,
y = rate,
group = group)) +
geom_ribbon(aes(ymin = lower.CL,
ymax = upper.CL,
fill = group),
alpha = .2) +
geom_line(aes(color = group,
linetype = group)) +
theme_bw() +
labs(x = "Four-Hour Intervals",
y = "Estimated Rate\nof Doses per 4 Hour Window",
color = "Dose",
fill = "Dose",
linetype = "Dose") +
scale_x_continuous(breaks = seq(from = 0, to = 12, by = 2)) +
theme(legend.position = "inside",
legend.position.inside = c(1, 1),
legend.justification = c(1.1, 1.1),
legend.background = element_rect(color = "black"),
legend.key.width = unit(1.5, "cm"))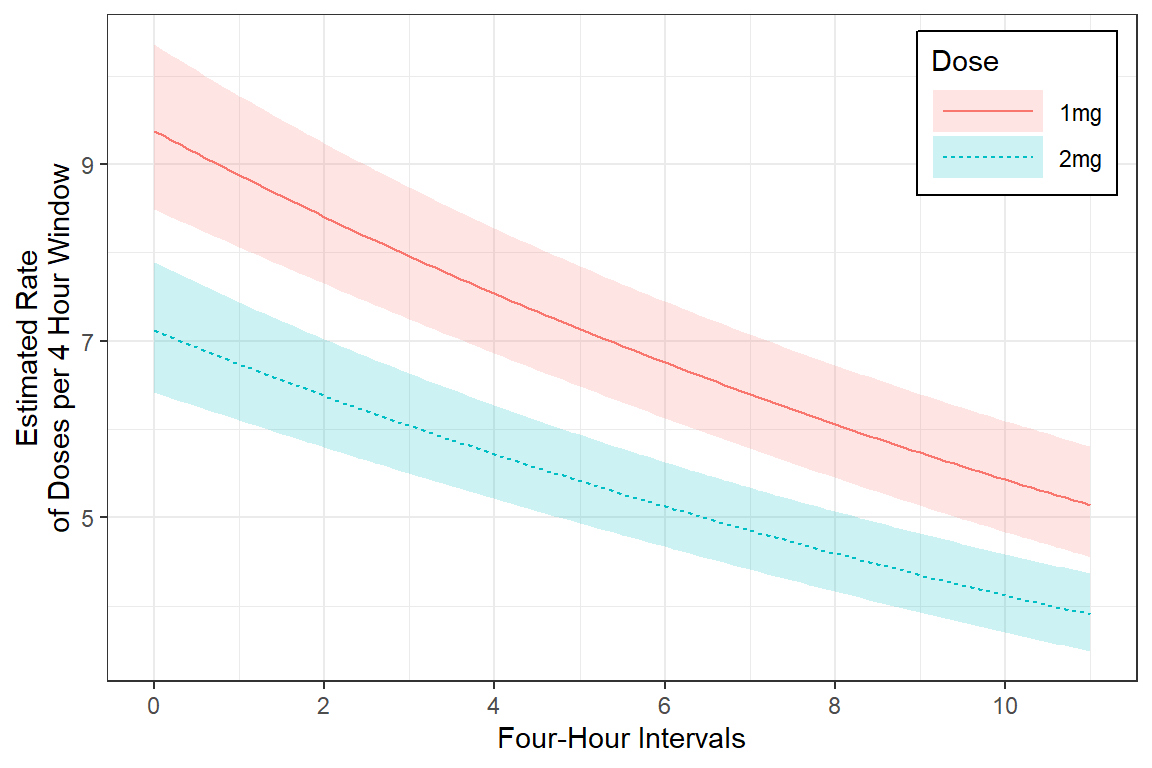
23.5 GLMM Analysis
23.5.1 RI: Random Intercepts Only
mod_glmer_ri <- lme4::glmer(count ~ group + I(time/4) + I((time/4)^2) + (1|id),
data = df_long,
family = poisson(link = "log"))
summary(mod_glmer_ri)Generalized linear mixed model fit by maximum likelihood (Laplace
Approximation) [glmerMod]
Family: poisson ( log )
Formula: count ~ group + I(time/4) + I((time/4)^2) + (1 | id)
Data: df_long
AIC BIC logLik -2*log(L) df.resid
4615 4638 -2303 4605 775
Scaled residuals:
Min 1Q Median 3Q Max
-3.505 -1.103 -0.174 0.825 8.581
Random effects:
Groups Name Variance Std.Dev.
id (Intercept) 0.246 0.496
Number of obs: 780, groups: id, 65
Fixed effects:
Estimate Std. Error z value Pr(>|z|)
(Intercept) 2.1347 0.0912 23.40 < 2e-16 ***
group2mg -0.2713 0.1276 -2.13 0.034 *
I(time/4) -0.2855 0.0593 -4.81 1.5e-06 ***
I((time/4)^2) 0.0256 0.0217 1.18 0.238
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Correlation of Fixed Effects:
(Intr) grp2mg I(t/4)
group2mg -0.641
I(time/4) -0.284 0.000
I((tm/4)^2) 0.231 0.000 -0.96023.5.2 RIAS: Random Intercepts and Slopes
mod_glmer_rias <- lme4::glmer(count ~ group + I(time/4) + I((time/4)^2) +
(I(time/4)|id),
data = df_long,
family = poisson(link = "log"))
summary(mod_glmer_rias)Generalized linear mixed model fit by maximum likelihood (Laplace
Approximation) [glmerMod]
Family: poisson ( log )
Formula: count ~ group + I(time/4) + I((time/4)^2) + (I(time/4) | id)
Data: df_long
AIC BIC logLik -2*log(L) df.resid
4440 4473 -2213 4426 773
Scaled residuals:
Min 1Q Median 3Q Max
-3.462 -0.965 -0.127 0.758 4.554
Random effects:
Groups Name Variance Std.Dev. Corr
id (Intercept) 0.300 0.548
I(time/4) 0.102 0.320 -0.45
Number of obs: 780, groups: id, 65
Fixed effects:
Estimate Std. Error z value Pr(>|z|)
(Intercept) 2.1060 0.0960 21.94 <2e-16 ***
group2mg -0.2481 0.1274 -1.95 0.0515 .
I(time/4) -0.2258 0.0726 -3.11 0.0019 **
I((time/4)^2) -0.0156 0.0222 -0.70 0.4837
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Correlation of Fixed Effects:
(Intr) grp2mg I(t/4)
group2mg -0.602
I(time/4) -0.412 -0.008
I((tm/4)^2) 0.226 0.000 -0.792Data: df_long
Models:
mod_glmer_ri: count ~ group + I(time/4) + I((time/4)^2) + (1 | id)
mod_glmer_rias: count ~ group + I(time/4) + I((time/4)^2) + (I(time/4) | id)
npar AIC BIC logLik -2*log(L) Chisq Df Pr(>Chisq)
mod_glmer_ri 5 4615 4638 -2303 4605
mod_glmer_rias 7 4440 4473 -2213 4426 179 2 <2e-16 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1apaSupp::tab_glmers(list("RI" = mod_glmer_ri,
"RIAS" = mod_glmer_rias),
caption = "GLMM: Main Effects Models")
| RI | RIAS | ||||
|---|---|---|---|---|---|---|
Variable | IRR | [95% CI] | p | IRR | [95% CI] | p |
(Intercept) | 8.45 | [7.07, 10.11] | < .001*** | 8.22 | [6.81, 9.92] | < .001*** |
group | ||||||
1mg | — | — | — | — | ||
2mg | 0.76 | [0.59, 0.98] | .034* | 0.78 | [0.61, 1.00] | .052 |
I(time/4) | 0.75 | [0.67, 0.84] | < .001*** | 0.80 | [0.69, 0.92] | .002** |
I((time/4)^2) | 1.03 | [0.98, 1.07] | .238 | 0.98 | [0.94, 1.03] | .484 |
id.var__(Intercept) | 0.25 | 0.30 | ||||
id.cov__(Intercept).I(time/4) | -0.08 | |||||
id.var__I(time/4) | 0.10 | |||||
AIC | 4,615.17 | 4,440.36 | ||||
BIC | 4,638.47 | 4,472.97 | ||||
Log-likelihood | -2,302.59 | -2,213.18 | ||||
Note. N = 780. CI = confidence interval. | ||||||
* p < .05. ** p < .01. *** p < .001. | ||||||
23.5.3 RAIS: Add Interaction
See the GLMM - Optimizers page for more information on convergence problems.
mod_glmer_rias2 <- lme4::glmer(count ~ group*I(time/4) + group*I((time/4)^2) +
( I(time/4)|id ),
data = df_long,
control = glmerControl(optimizer ="Nelder_Mead"),
family = poisson(link = "log")) Data: df_long
Models:
mod_glmer_rias: count ~ group + I(time/4) + I((time/4)^2) + (I(time/4) | id)
mod_glmer_rias2: count ~ group * I(time/4) + group * I((time/4)^2) + (I(time/4) | id)
npar AIC BIC logLik -2*log(L) Chisq Df Pr(>Chisq)
mod_glmer_rias 7 4440 4473 -2213 4426
mod_glmer_rias2 9 4442 4484 -2212 4424 1.93 2 0.38apaSupp::tab_glmers(list("Main Effects" = mod_glmer_rias,
"interactions" = mod_glmer_rias2),
caption = "GLMM: Main Effects vs. Interaction")
| Main Effects | interactions | ||||
|---|---|---|---|---|---|---|
Variable | IRR | [95% CI] | p | IRR | [95% CI] | p |
(Intercept) | 8.22 | [6.81, 9.92] | < .001*** | 7.84 | [6.40, 9.60] | < .001*** |
group | ||||||
1mg | — | — | — | — | ||
2mg | 0.78 | [0.61, 1.00] | .052 | 0.87 | [0.64, 1.18] | .366 |
I(time/4) | 0.80 | [0.69, 0.92] | .002** | 0.87 | [0.72, 1.04] | .133 |
I((time/4)^2) | 0.98 | [0.94, 1.03] | .484 | 0.97 | [0.91, 1.02] | .211 |
group * I(time/4) | ||||||
2mg * I(time/4) | 0.81 | [0.61, 1.09] | .162 | |||
group * I((time/4)^2) | ||||||
2mg * I((time/4)^2) | 1.05 | [0.96, 1.15] | .253 | |||
id.var__(Intercept) | 0.30 | 0.30 | ||||
id.cov__(Intercept).I(time/4) | -0.08 | -0.08 | ||||
id.var__I(time/4) | 0.10 | 0.10 | ||||
AIC | 4,440.36 | 4,442.42 | ||||
BIC | 4,472.97 | 4,484.36 | ||||
Log-likelihood | -2,213.18 | -2,212.21 | ||||
Note. N = 780. CI = confidence interval. | ||||||
* p < .05. ** p < .01. *** p < .001. | ||||||
23.5.4 RIAS: Simplify
mod_glmer_rias3 <- lme4::glmer(count ~ group + I(time/4) +
( I(time/4)|id ),
data = df_long,
control = glmerControl(optimizer ="Nelder_Mead"),
family = poisson(link = "log")) Data: df_long
Models:
mod_glmer_rias3: count ~ group + I(time/4) + (I(time/4) | id)
mod_glmer_rias: count ~ group + I(time/4) + I((time/4)^2) + (I(time/4) | id)
npar AIC BIC logLik -2*log(L) Chisq Df Pr(>Chisq)
mod_glmer_rias3 6 4439 4467 -2213 4427
mod_glmer_rias 7 4440 4473 -2213 4426 0.49 1 0.48apaSupp::tab_glmers(list("Quadratic" = mod_glmer_rias,
"Linear" = mod_glmer_rias3),
caption = "GLMM: Nature of Time")
| Quadratic | Linear | ||||
|---|---|---|---|---|---|---|
Variable | IRR | [95% CI] | p | IRR | [95% CI] | p |
(Intercept) | 8.22 | [6.81, 9.92] | < .001*** | 8.34 | [6.95, 10.01] | < .001*** |
group | ||||||
1mg | — | — | — | — | ||
2mg | 0.78 | [0.61, 1.00] | .052 | 0.78 | [0.61, 1.00] | .051 |
I(time/4) | 0.80 | [0.69, 0.92] | .002** | 0.77 | [0.70, 0.84] | < .001*** |
I((time/4)^2) | 0.98 | [0.94, 1.03] | .484 | |||
id.var__(Intercept) | 0.30 | 0.30 | ||||
id.cov__(Intercept).I(time/4) | -0.08 | -0.08 | ||||
id.var__I(time/4) | 0.10 | 0.10 | ||||
AIC | 4,440.36 | 4,438.85 | ||||
BIC | 4,472.97 | 4,466.80 | ||||
Log-likelihood | -2,213.18 | -2,213.42 | ||||
Note. N = 780. CI = confidence interval. | ||||||
* p < .05. ** p < .01. *** p < .001. | ||||||
23.5.5 Visualize Best Model
23.5.5.1 Model Parameter Table
Incident Rate Ratio | Log Scale | ||||||
|---|---|---|---|---|---|---|---|
Variable | IRR | [95% CI] | b | (SE) | p | ||
(Intercept) | 8.34 | [6.95, 10.01] | |||||
group | |||||||
1mg | — | — | — | — | |||
2mg | 0.78 | [0.61, 1.00] | -0.25 | (0.13) | .051 | ||
I(time/4) | 0.77 | [0.70, 0.84] | -0.27 | (0.04) | < .001*** | ||
id.var__(Intercept) | 0.30 | ||||||
id.cov__(Intercept).I(time/4) | -0.08 | ||||||
id.var__I(time/4) | 0.10 | ||||||
Note. p = significance from Wald t-test for parameter estimate using Satterthwaite's method for degrees of freedom. | |||||||
* p < .05. ** p < .01. *** p < .001. | |||||||
23.5.5.2 Estimated Marginal Means
mod_glmer_rias3 %>%
emmeans::emmeans(pairwise ~group|time,
at = list(time = c(0, 5, 10)),
type = "response")$emmeans
time = 0:
group rate SE df asymp.LCL asymp.UCL
1mg 8.34 0.778 Inf 6.95 10.01
2mg 6.51 0.658 Inf 5.34 7.93
time = 5:
group rate SE df asymp.LCL asymp.UCL
1mg 5.98 0.526 Inf 5.03 7.11
2mg 4.67 0.445 Inf 3.87 5.62
time = 10:
group rate SE df asymp.LCL asymp.UCL
1mg 4.29 0.485 Inf 3.44 5.35
2mg 3.35 0.395 Inf 2.65 4.22
Confidence level used: 0.95
Intervals are back-transformed from the log scale
$contrasts
time = 0:
contrast ratio SE df null z.ratio p.value
1mg / 2mg 1.28 0.163 Inf 1 1.948 0.0514
time = 5:
contrast ratio SE df null z.ratio p.value
1mg / 2mg 1.28 0.163 Inf 1 1.948 0.0514
time = 10:
contrast ratio SE df null z.ratio p.value
1mg / 2mg 1.28 0.163 Inf 1 1.948 0.0514
Tests are performed on the log scale 23.5.5.3 Plot Estimated Marginal Means
interactions::interact_plot(model = mod_glmer_rias3,
pred = time,
modx = group,
interval = TRUE,
int.width = .685)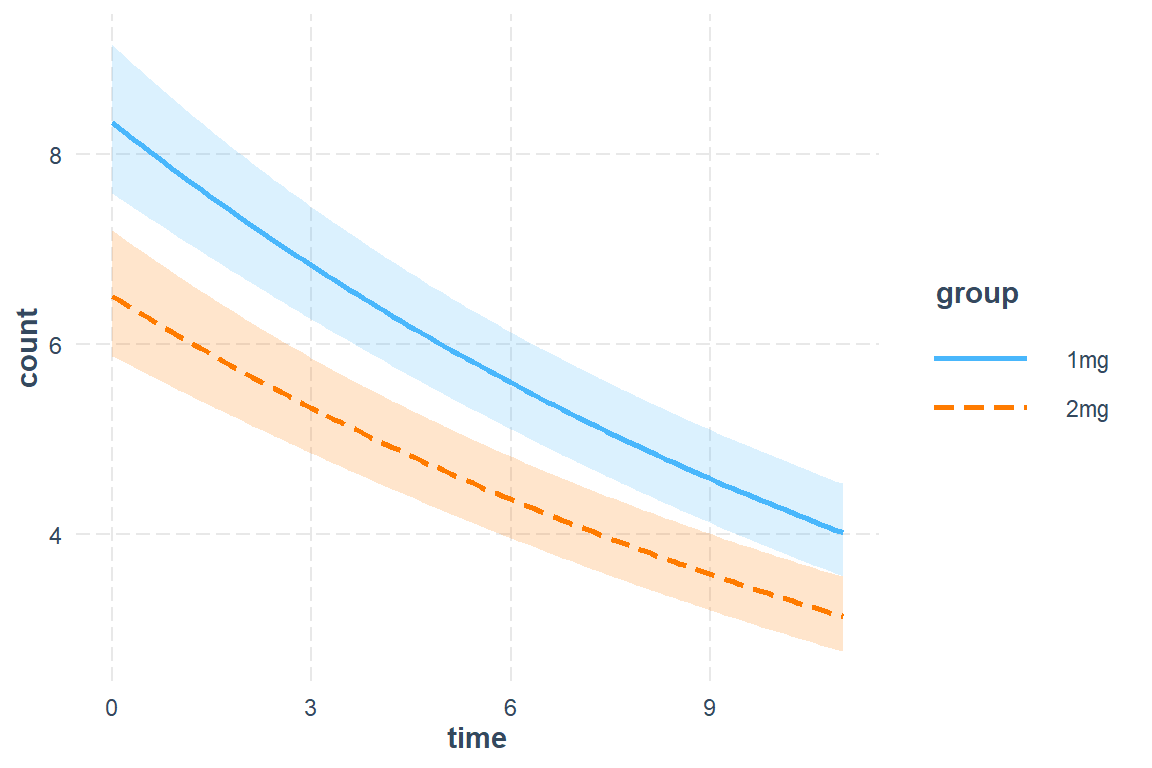
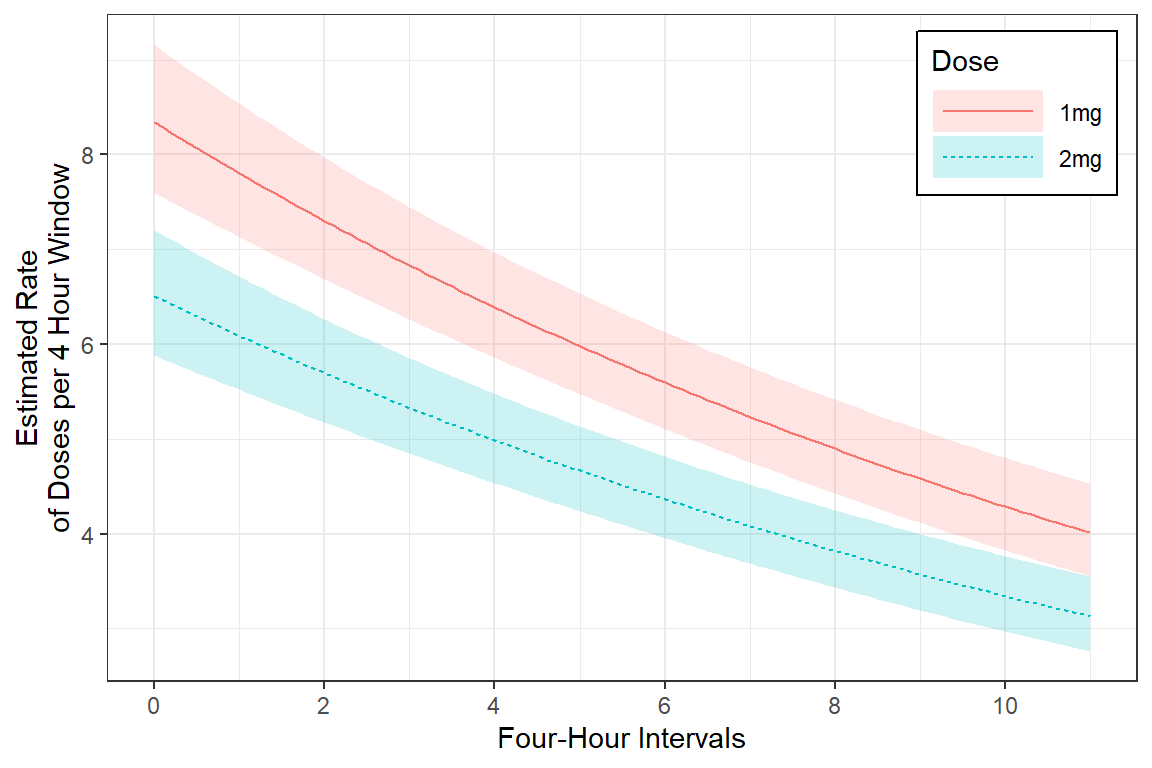
mod_glmer_rias3 %>%
emmeans::emmeans(~group|time,
at = list(time = seq(from = 0, to = 11, by = .1)),
type = "response",
level = .685) %>%
data.frame() %>%
ggplot(aes(x = time,
y = rate,
group = group)) +
geom_ribbon(aes(ymin = asymp.LCL,
ymax = asymp.UCL,
fill = group),
alpha = .2) +
geom_line(aes(color = group,
linetype = group)) +
theme_bw() +
labs(x = "Four-Hour Intervals",
y = "Estimated Rate\nof Doses per 4 Hour Window",
color = "Dose",
fill = "Dose",
linetype = "Dose") +
scale_x_continuous(breaks = seq(from = 0, to = 12, by = 2)) +
theme(legend.position = "inside",
legend.position.inside = c(1, 1),
legend.justification = c(1.1, 1.1),
legend.background = element_rect(color = "black"),
legend.key.width = unit(1.5, "cm"))